Post-translational modification (PTM) refers to the covalent and generally enzymatic modification of proteins during or after protein biosynthesis. Proteins are synthesized by ribosomes translating mRNA into polypeptide chains, which may then undergo PTM to form the mature protein product. PTMs are important components in cell signaling.
Post-translational modifications can occur on the amino acid side chains or at the protein's C- or N- termini. They can extend the chemical repertoire of the 20 standard amino acids by modifying an existing functional group or introducing a new one such as phosphate. Phosphorylation is a very common mechanism for regulating the activity of enzymes and is the most common post-translational modification. Many eukaryotic proteins also have carbohydrate molecules attached to them in a process called glycosylation, which can promote protein folding and improve stability as well as serving regulatory functions. Attachment of lipid molecules, known as lipidation, often targets a protein or part of a protein attached to the cell membrane.
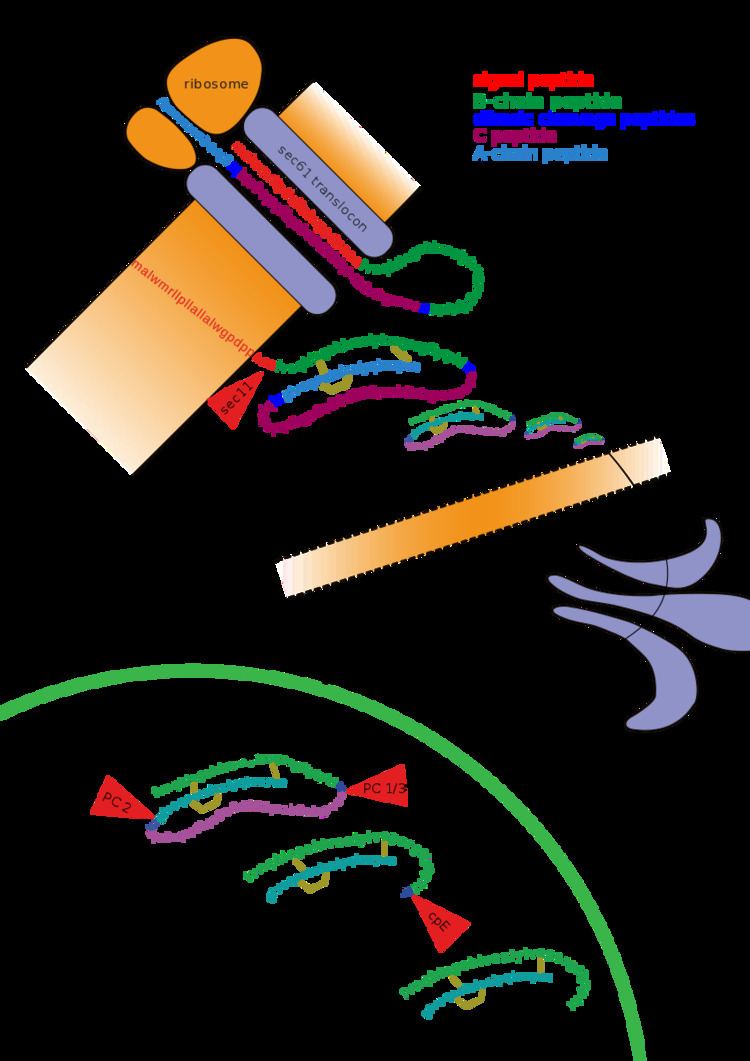
Other forms of post-translational modification consist of cleaving peptide bonds, as in processing a propeptide to a mature form or removing the initiator methionine residue. The formation of disulfide bonds from cysteine residues may also be referred to as a post-translational modification. For instance, the peptide hormone insulin is cut twice after disulfide bonds are formed, and a propeptide is removed from the middle of the chain; the resulting protein consists of two polypeptide chains connected by disulfide bonds.
Some types of post-translational modification are consequences of oxidative stress. Carbonylation is one example that targets the modified protein for degradation and can result in the formation of protein aggregates. Specific amino acid modifications can be used as biomarkers indicating oxidative damage.
Sites that often undergo post-translational modification are those that have a functional group that can serve as a nucleophile in the reaction: the hydroxyl groups of serine, threonine, and tyrosine; the amine forms of lysine, arginine, and histidine; the thiolate anion of cysteine; the carboxylates of aspartate and glutamate; and the N- and C-termini. In addition, although the amide of asparagine is a weak nucleophile, it can serve as an attachment point for glycans. Rarer modifications can occur at oxidized methionines and at some methylenes in side chains.
Post-translational modification of proteins can be experimentally detected by a variety of techniques, including mass spectrometry, Eastern blotting, and Western blotting.
myristoylation (a type of acylation), attachment of myristate, a C14 saturated acidpalmitoylation (a type of acylation), attachment of palmitate, a C16 saturated acidisoprenylation or prenylation, the addition of an isoprenoid group (e.g. farnesol and geranylgeraniol)farnesylationgeranylgeranylationglypiation, glycosylphosphatidylinositol (GPI) anchor formation via an amide bond to C-terminal taillipoylation (a type of acylation), attachment of a lipoate (C8) functional groupflavin moiety (FMN or FAD) may be covalently attachedheme C attachment via thioether bonds with cysteinesphosphopantetheinylation, the addition of a 4'-phosphopantetheinyl moiety from coenzyme A, as in fatty acid, polyketide, non-ribosomal peptide and leucine biosynthesisretinylidene Schiff base formationdiphthamide formation (on a histidine found in eEF2)ethanolamine phosphoglycerol attachment (on glutamate found in eEF1α)hypusine formation (on conserved lysine of eIF5A (eukaryotic) and aIF5A (archaeal))acylation, e.g. O-acylation (esters), N-acylation (amides), S-acylation (thioesters)acetylation, the addition of an acetyl group, either at the N-terminus of the protein or at lysine residues. See also histone acetylation. The reverse is called deacetylation.formylationalkylation, the addition of an alkyl group, e.g. methyl, ethylmethylation the addition of a methyl group, usually at lysine or arginine residues. The reverse is called demethylation.amide bond formationamidation at C-terminusamino acid additionarginylation, a tRNA-mediation additionpolyglutamylation, covalent linkage of glutamic acid residues to the N-terminus of tubulin and some other proteins. (See tubulin polyglutamylase)polyglycylation, covalent linkage of one to more than 40 glycine residues to the tubulin C-terminal tailbutyrylationgamma-carboxylation dependent on Vitamin Kglycosylation, the addition of a glycosyl group to either arginine, asparagine, cysteine, hydroxylysine, serine, threonine, tyrosine, or tryptophan resulting in a glycoprotein. Distinct from glycation, which is regarded as a nonenzymatic attachment of sugars.polysialylation, addition of polysialic acid, PSA, to NCAMmalonylationhydroxylation: addition of an oxygen atom to the side-chain of a Pro or Lys residueiodination: addition of an iodine atom to the aromatic ring of a tyrosine residue (e.g. in thyroglobulin)nucleotide addition such as ADP-ribosylationphosphate ester (O-linked) or phosphoramidate (N-linked) formationphosphorylation, the addition of a phosphate group, usually to serine, threonine, and tyrosine (O-linked), or histidine (N-linked)adenylylation, the addition of an adenylyl moiety, usually to tyrosine (O-linked), or histidine and lysine (N-linked)propionylationpyroglutamate formationS-glutathionylationS-nitrosylationS-sulfenylation (aka S-sulphenylation), reversible covalent addition of one oxygen atom to the thiol group of a cysteine residueS-sulfinylation, normally irreversible covalent addition of two oxygen atoms to the thiol group of a cysteine residueS-sulfonylation, normally irreversible covalent addition of three oxygen atoms to the thiol group of a cysteine residue, resulting in the formation of a cysteic acid residuesuccinylation addition of a succinyl group to lysinesulfation, the addition of a sulfate group to a tyrosine.glycation, the addition of a sugar molecule to a protein without the controlling action of an enzyme.carbamylation the addition of Isocyanic acid to a protein's N-terminus or the side-chain of Lys.carbonylation the addition of carbon monoxide to other organic/inorganic compounds.biotinylation: covalent attachment of a biotin moiety using a biotinylation reagent, typically for the purpose of labeling a protein.carbamylation: the addition of Isocyanic acid to a protein's N-terminus or the side-chain of Lys or Cys residues, typically resulting from exposure to urea solutions.oxidation: addition of one or more Oxygen atoms to a susceptible side-chain, principally of Met, Trp, His or Cys residues. Formation of disulfide bonds between Cys residues.pegylation: covalent attachment of polyethylene glycol (PEG) using a pegylation reagent, typically to the N-terminus or the side-chains of Lys residues. Pegylation is used to improve the efficacy of protein pharmaceuticals.ISGylation, the covalent linkage to the ISG15 protein (Interferon-Stimulated Gene 15)SUMOylation, the covalent linkage to the SUMO protein (Small Ubiquitin-related MOdifier)ubiquitination, the covalent linkage to the protein ubiquitin.Neddylation, the covalent linkage to NeddPupylation, the covalent linkage to the Prokaryotic ubiquitin-like proteincitrullination, or deimination, the conversion of arginine to citrulline deamidation, the conversion of glutamine to glutamic acid or asparagine to aspartic acideliminylation, the conversion to an alkene by beta-elimination of phosphothreonine and phosphoserine, or dehydration of threonine and serine disulfide bridges, the covalent linkage of two cysteine amino acidsproteolytic cleavage, cleavage of a protein at a peptide bondisoaspartate formation, via the cyclisation of asparagine or aspartic acid amino-acid residuesracemizationof serine by protein-serine epimeraseof alanine in dermorphin, a frog opioid peptideof methionine in deltorphin, also a frog opioid peptideprotein splicing, self-catalytic removal of inteins analogous to mRNA processingIn 2011, statistics of each post-translational modification experimentally and putatively detected have been compiled using proteome-wide information from the Swiss-Prot database. The 10 most common experimentally found modifications were as follows:
More details can be found at http://selene.princeton.edu/PTMCuration/.
Some common post-translational modifications to specific amino-acid residues are shown below. Modifications occur on the side-chain unless indicated otherwise.
Cleavage and formation of disulfide bridges during the production of insulinPTM of histones as regulation of transcription: RNA polymerase control by chromatin structurePTM of RNA polymerase II as regulation of transcriptionCleavage of polypeptide chains as crucial for lectin specificity