 | ||
The Walker A and Walker B motifs are protein sequence motifs, now known to have highly conserved three-dimensional structures. These were first reported in ATP-binding proteins by Walker and co-workers in 1982.
Contents
Walker A motif
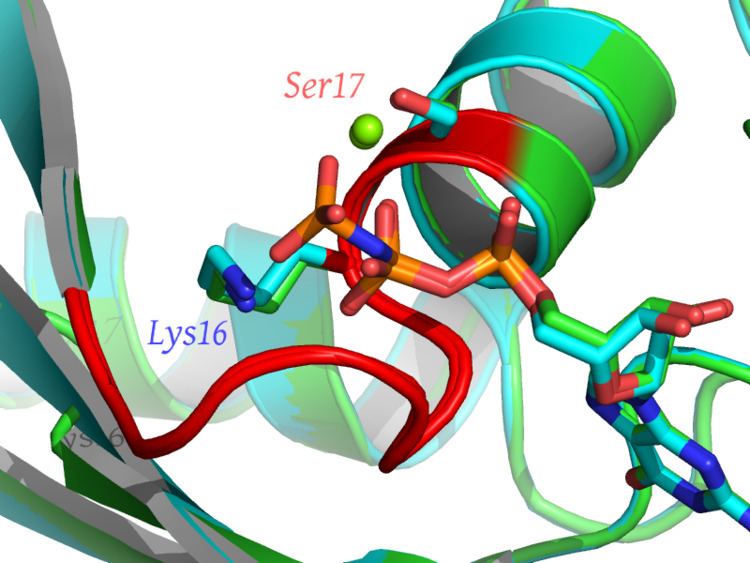
Walker A motif, also known as the Walker loop or P-loop (phosphate-binding loop) is a motif in proteins that is associated with phosphate binding. The motif has the pattern G-x(4)-GK-[TS], where G, K, T and S denote glycine, lysine, threonine and serine residues respectively, and x denotes any amino acid. It is present in many ATP or GTP utilizing proteins; it is the β phosphate of the nucleotide that is bound. The lysine (K) residue in the Walker A motif, together with the main chain NH atoms, are crucial for nucleotide-binding. It is a glycine-rich loop preceded by a beta strand and followed by an alpha helix; these features are typically part of an α/β domain with four strands sandwiched between two helices on each side. The phosphate groups of the nucleotide are also coordinated to a divalent cation such as a magnesium, calcium, or manganese(II) ion.
Apart from the conserved lysine, a feature of the P-loop used in phosphate binding is a compound LRLR nest comprising the four residues xxGK, as above, whose mainchain atoms form a phosphate-sized concavity with the NH groups pointing inwards. The hexapeptide SGAGKT has been shown to bind inorganic phosphate strongly; since such a short peptide does not form an alpha helix, this suggests that it is the nest, rather than being at the N-terminus of a helix, that is the main phosphate binding feature.
The Walker A motif is best known for its presence in ATP- and GTP-binding proteins, and is also found in a variety of proteins with phosphorylated substrates. These include ATP synthase (α and β subunits), myosin, transducin, helicases, kinases, AAA proteins, G-proteins, RecA, protein tyrosine phosphatases (see below) and pyridoxal phosphate utilizing enzymes such as cysteine synthase.
Upon nucleotide hydrolysis the loop does not significantly change conformation, but stays bound to the remaining phosphate groups. Walker motif A-binding has been shown to cause structural changes in the bound nucleotide, along the line of the induced fit model of enzyme binding.
PTPs (protein tyrosine phosphatases) that catalyse the hydrolysis of an inorganic phosphate from a phosphotyrosine residue (the reverse of a tyrosine kinase reaction) contains a motif which folds into a P-loop-like structure with an arginine in the place of the conserved lysine. The conserved sequence of this motif is C-x(5)-R-[ST], where C and R denote cysteine and arginine residues respectively.
A-loop
The A-loop (Aromatic residue interacting with the adenine ring of ATP) refers to conserved aromatic amino acids, essential for ATP-binding, found about 25 amino acids upstream of the Walker A motif in a subset of P-loop proteins.
Walker B motif
Walker B motif is a motif in most P-loop proteins situated well downstream of the A-motif. Walker et al reported the consensus sequence of this motif to be [RK]-x(4)-G-x(4)-LhhhhD, where R, K, G, L and D denote arginine, lysine, glycine, leucine and aspartic acid residues respectively, x represents any of the 20 standard amino acids and h denotes a hydrophobic amino acid. The consensus sequence of this motif was more recently reported by Hanson and Whiteheart (2005) to be hhhhDE, where E denotes a glutamate residue. The aspartate and glutamate also form a part of the DEAD/DEAH motifs found in helicases. The aspartate residue co-ordinates magnesium ions, and the glutamate is essential for ATP hydrolysis. There is considerable variability in the sequence of this motif, with the only invariant features being a negatively charged residue following a stretch of bulky, hydrophobic amino acids.
