Symbol IRES_HepA Rfam RF00228 | Alt. Symbols HepA_IRES GO 0043022 | |
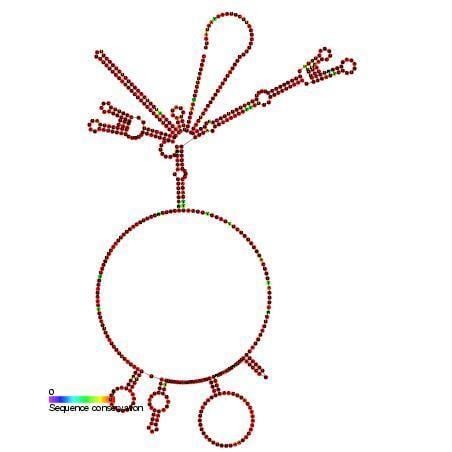 | ||
This family represents the internal ribosome entry site (IRES) of the hepatitis A virus. HAV IRES is a 450 nucleotide long sequence located in the 735 nt long 5’ UTR (untranslated region) of Hepatitis A viral RNA genome. IRES elements allow cap and end-independent translation of mRNA in the host cell. The IRES achieves this by mediating the internal initiation of translation by recruiting a ribosomal 40S pre-initiation complex directly to the initiation codon and eliminates the requirement for eukaryotic initiation factor, eIF4F.
Contents
History and Background
The IRES was first discovered in the RNA genome of picornaviridae by Norman Sonenberg in 1988. Initially, the identified 5’ UTR of poliovirus (PV) that directed internal initiation for protein synthesis was termed ribosome landing pad (RLP). This term was substituted with internal ribosome entry site soon after, which is also more commonly used today. In 1988, only encephalomycoarditis virus (EMCV) and PV were being used to demonstrate IRES ability. Research identifying the IRES in HAV was done in 1993 by Michael J. Glass, Xi-Yu Jia, and Donald F. Summers. Their research indicated the IRES of HAV was located downstream of nucleotide 45 and included up to nucleotide 734. Cap-independent internal initiation of protein synthesis differs from normal cellular cap-dependent translation initiation. eIF4 (eukaryotic initiation factor 4) as a whole complex made up of eIF4A, eIF4B, eIF4E, and eIF4G. eIF4F is used to refer to the complex that comprises eIF4E, 4G, and 4A. In cap-dependent translation initiation, the eIF4F binds to the m7G cap on the 5’ UTR end, recruiting the 40S small ribosomal subunit and scanning downstream for the AUG start codon. Cap-independent internal initiation has shown to interfere with either parts of eIF4 or the entire complex.
Function
Uncoated in the cytoplasm, HAV genome initiates translation independent of a 5’ cap in order to synthesize its viral proteins. The secondary structure of the HAV IRES is both necessary and sufficient for the genome to recruit a ribosome and initiate translation. A host cell ribosome recognizes the IRES and will directly enter at the sequence rather than scan from the 5’ end. The HAV genome does not encode for proteins that have host protein shut off abilities. Therefore, the HAV IRES must compete with host cell m7G capped mRNA. Unfortunately, HAV IRES initiation of translation is not as efficient as a typical host cell m7G cap. Although HAV IRES structure has affinity for eIF4F, its affinity is not nearly as high as the host cell’s capped mRNAs. This results in a longer period required until maximum shedding of the virus is reached. This also results in cell death only occurring from host immune responses rather than lysis.
Suppression of IRES
Because eIF4F plays an essential role for HAV IRES initiation, it is a target for suppression of IRES. Cleavage of eIF4G, a protein scaffold of eIF4F, by sequence specific proteases 2A protease or L-protease, will result in highly inhibited HAV IRES activity. These proteases are encoded by other members of the picornaviridae family. Foot-and-mouth disease virus (FMDV), for example, encodes these proteases to inhibit cellular mRNA translation while allowing for viral RNA to be translated. The requirement of an intact eIF4G for IRES initiation is specific to HAV IRES among other picornaviruses. eIF4E-binding protein I (4E-BP1) will also interfere with the eIF4G protein. 4E-BP1 functions by sequestering eIF4E which, thereby, inhibiting its association with eIF4G and resulting in HAV IRES inactivation (1). Another method to inactivate eIF4F activity is through the effects of the m7GpppG cap analogue, which targets eIF4E and is then able to prevent its association with capped 5’ ends of mRNAs. The exact mechanism in which this cap analogue interferes with IRES is not clear but it is suggested that the binding of this analogue to eIF4E results in a conformational change of eIF4G which is what disrupts eIF4G’s normal function.
glyceraldehyde-3-phosphate-dehydrogenase (GAPDH)
Glyceraldehyde-3-phosphate-dehydrogenase (GAPDH) is a cellular enzyme typically involved in glycolysis. GAPDH is known to bind to the overlapping sites within the stem-loop IIIa within the HAV IRES. The stem-loop IIIa contains a UU nucleotide deletion inside of a 5 nucleotide sequence which enhances the IRES activity. GAPDH effectively binding to this region will destabilize the secondary structure that the IRES forms, suppressing the IRES’s ability to perform the cap-independent translation.
La protein suppression
A host cell protein found widely and exclusively in eukaryotic cells, La protein binds directly to specific regions on the HAV IRES during mRNA translation as well as RNA replication. In a 2008 study, cytoplasmic La was observed to reduce HAV IRES initiation. However, in 2014, a more recent study demonstrated successful inhibition (in vivo) of La protein as a proposed method for inhibiting the HAV IRES translation and replication, which means that it more than likely plays an integral role in the HAV translation and replication.
Amantadine
Amantadine, a tricyclic symmetric amine, is a proven suppressor that specifically inhibits the HAV IRES dependent translation of HAV RNA. A 2005 experiment showed amantadine suppressed HAV IRES translation and did not trigger an interferon response, which indicates promising antiviral usage of amantadine. For influenza A virus, its primary method of action as an antiviral is to prevent the uncoating of viral genome which inhibits the HAV IRES- mediated translation and replication. Amantadine’s effectiveness stems from the IRES location on the 5’NTR region which has a high affinity for antivirals making it an effective target. It was also revealed that the M2 protein of influenza A virus could be another viable target for the potential antiviral .
Homologs of IRES across picornaviridae (family)
All picornaviruses have been found to contain IRES. There are four classes of IRES within the picornaviridae family, ranging from 270-450 nt. Among picornaviruses, many 5’ UTRs also contain additional structural elements upstream, which can assist the viral genome in replication. Many picornavirus IRES also permit many viruses to block cap-dependent initiation, which results in shutting off host cell protein synthesis. The four class are Entero-/rhinovirus IRES, Cardio-/aphthovirus IRES, HAV IRES, HCV-like picornavirus IRES. These IRES are categorized by their nucleotide sequences but share structural similarity because it is the RNA structure that has the ability to internally recruit translational machinery. Entero-/rhinovirus IRES elements share some structural motifs with HAV IRES. HAV IRES, entero-/rhinovirus and cardio-/apthovirus IRES are all approximately 450 nt but differ greatly in their structures. A cardiovirus, EMCV, and an apthovirus, foot-and-mouth disease virus (FMDV) share about 50% identical IRES elements. HCV-like picornavirus IRES incorporates the most difference in IRES elements from the other three classes. There is a wide variety of picornaviridae viruses that have highly conserved HCV-like IRES elements, some of which are predicted to still be identified. It is important to note that HAV IRES activity differs from the other three classes in its specific requirement for an intact eIF4G. Other picornaviruses encode for proteins that will cleave the eIF4G for enhanced IRES activity.
