 | ||
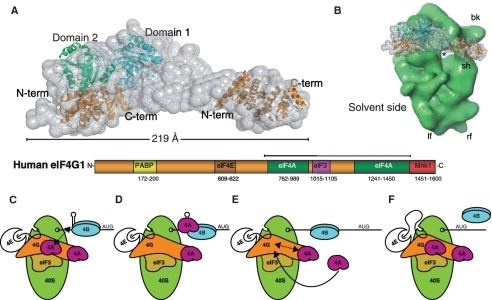
Eukaryotic translation initiation factor 4 G (eIF4G) is a protein involved in eukaryotic translation initiation, and is a component of the eIF4F cap binding complex. Orthologs of eIF4G have been studied in multiple species, including humans, yeast, and wheat. However, eIF4G is exclusively found in domain Eukarya, and not in domains Bacteria or Archaea, which do not have capped mRNA. As such, eIF4G structure and function may vary between species, although the human eIF4G 1 has been the focus of extensive studies.
Contents
- History
- eIF4G Binding Partners
- eIF4G in Translation Initiation
- eIF4G in Disease
- Role in Aging
- Importance in Virology
- References
Across species, eIF4G strongly associates with the protein that directly binds the mRNA cap: eIF4E. The eIF4F complex contain eIF4G, eIF4E, and the RNA helicase protein eIF4A).
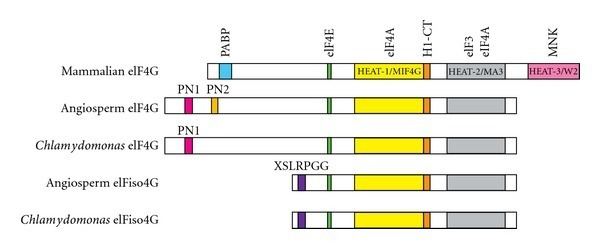
Within the cell eIF4G is found primarily in the cytoplasm, usually bound to eIF4E; however, it is also found in the nucleus where its function is unknown. It may also to have a role in nonsense-mediated decay.
History
eIF4G stands for eukaryotic initiation factor 4 gamma (typically gamma is now replaced by G in the literature). It was initially isolated by fractionation, found present in fraction 4 gamma, and was involved in eukaryotic translation initiation.
eIF4G Binding Partners
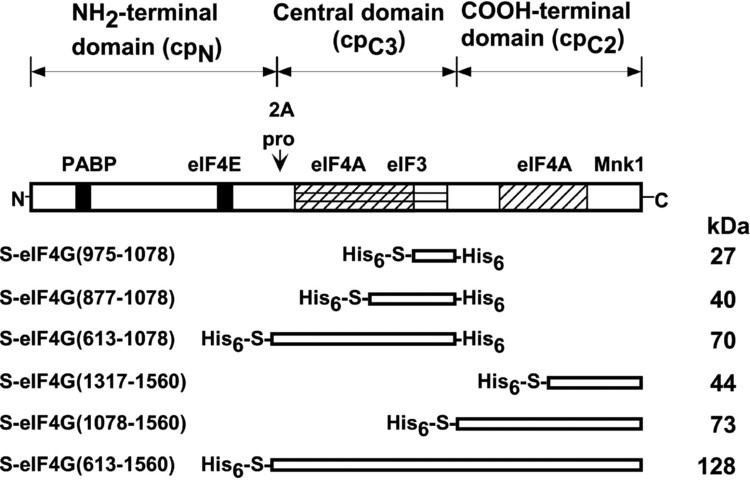
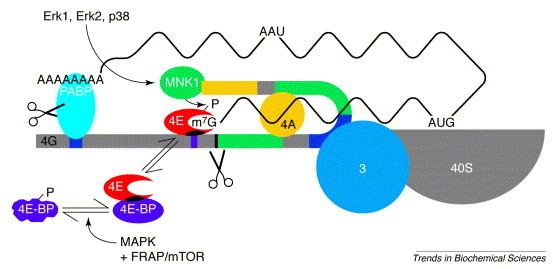
eIF4G has been found to associate with myriad other proteins besides those of the eIF4F complex, including MNK-1, CBP80, CBP20, PABP, and eIF3. eIF4G also directly binds mRNA and has multiple positively charged regions for this function. Several IRESs also bind eIF4G directly, as do BTE CITEs.
eIF4G in Translation Initiation
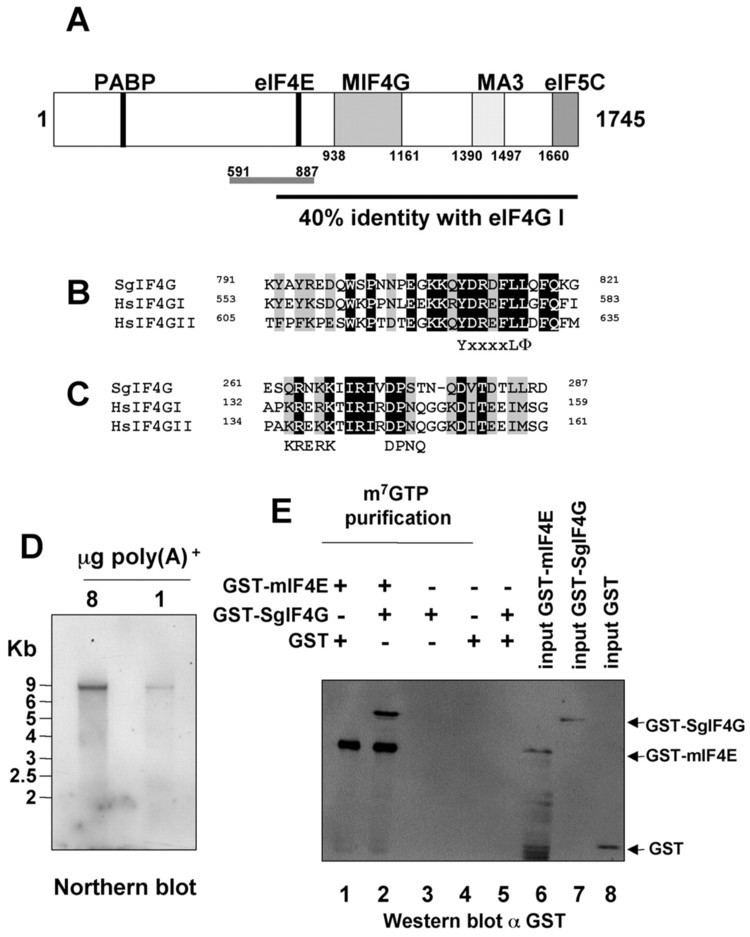
eIF4G is an important scaffold for the eIF4F complex and aids in recruiting the 40S ribosomal subunit to mRNA.
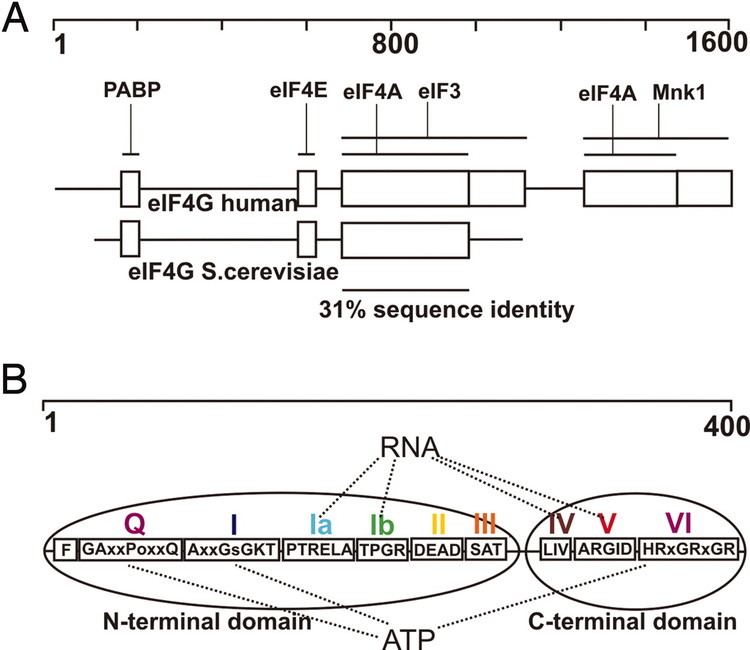
There are 3 mechanisms that the 40S ribosome can come to recognize the start codon: scanning, internal entry, and shunting. Scanning is where the 40S ribosome slides along the RNA until recognizing a start site (typically an AUG sequence in "good context"). Internal entry is where the 40S ribosome does not start from the beginning (5' end) of the mRNA but instead starts from somewhere in the middle. Shunting is where after the 40S ribosome starts sliding along the mRNA it "jumps" or skips large sections; the mechanism for this is still unclear. eIF4G is required for most types of initiation, exception special cases such as internal initiation by HCV IRES or Cripavirus internal ribosome entry site (IRES).
eIF4G in Disease
eIF4G has been implicated in breast cancer. It appears in increased levels in certain types of breast cancer (and others) and increases IRES containing mRNA production; these mRNAs produce hypoxia and stress related proteins that encourage blood vessel invasion (which is important for cancer cells—requiring large amounts of nutrients to divide quickly and form sizable tumors).
Role in Aging
Regulation of translation initiation by eIF4G is vital for protein synthesis in developing organisms, for example yeast and nematodes. Deletion of eIF4G is lethal in yeast. In the round worm, C. elegans, knockout of eIF4G leads to animals that cannot develop past the early larval stage (L2) of development. The critical role of eIF4G in development appears to be reversed in adulthood, when eIF4G dysregulation negatively impacts lifespan and increases susceptibility to certain aging-related diseases (see eIF4G in diseases above). Inhibiting eIF4G during adulthood in C. elegans drastically extends lifespan, comparable to the lifespan increase exhibited during dietary restriction. In addition, inhibiting eIF4G reduces overall protein translation, while preferentially translating mRNA of genes important for responding to stress and against those associated with growth and reproduction. Thus eIF4G appears to control differential mRNA translation during periods or growth and stress, which may ultimately lead to age related decline.
Importance in Virology
As previously mentioned, eIF4G is bound by certain IRESs, which were initially discovered in viruses. Some viral IRESs directly bind eIF4G, and co-opt it for gaining access to the ribosome. Some cellular mRNAs also contain IRESs (including eIF4G itself).
Some viral proteases cleave off part of eIF4G, that contains the eIF4E binding region. This has the effect of preventing most cellular mRNAs from binding eIF4G; however, a few cellular mRNAs with IRESs still translate under these conditions.
One example of a, eIF4G binding site in a viral IRES's is in the EMCV IRES (nucleotides 746-949).
