Group Group I (dsDNA) Family Myoviridae Genus T4virus Higher classification T4virus | Order Caudovirales Subfamily Tevenvirinae Scientific name Enterobacteria phage T4 Rank Species | |
 | ||
Similar T7 phage, Escherichia, Lambda phage, M13 bacteriophage, Eukaryote | ||
Enterobacteria phage T4 is a bacteriophage that infects Escherichia coli bacteria. The T4 phage is a member of the T-even phages, a group including enterobacteriophages T2 and T6. T4 is capable of undergoing only a lytic lifecycle and not the lysogenic lifecycle.
Contents
- T4 Phage attacking Ecoli
- Genome and structure
- Translation
- Virus particle structure
- Infection process
- Reproduction
- Replication and packaging
- Multiplicity reactivation
- History
- References
T4 Phage attacking E.coli
Genome and structure
The T4 phage's double-stranded DNA genome is about 169 kbp long and encodes 289 proteins. The T4 genome is terminally redundant and is first replicated as a unit, then several genomic units are recombined end-to-end to form a concatemer. When packaged, the concatemer is cut at unspecific positions of the same length, leading to several genomes that represent circular permutations of the original. The T4 genome bears eukaryote-like intron sequences.
Translation
The Shine-Dalgarno sequence GAGG dominates in bacteriophage T4 early genes, whereas the sequence GGAG is a target for the T4 endonuclease RegB that initiates the early mRNA degradation.
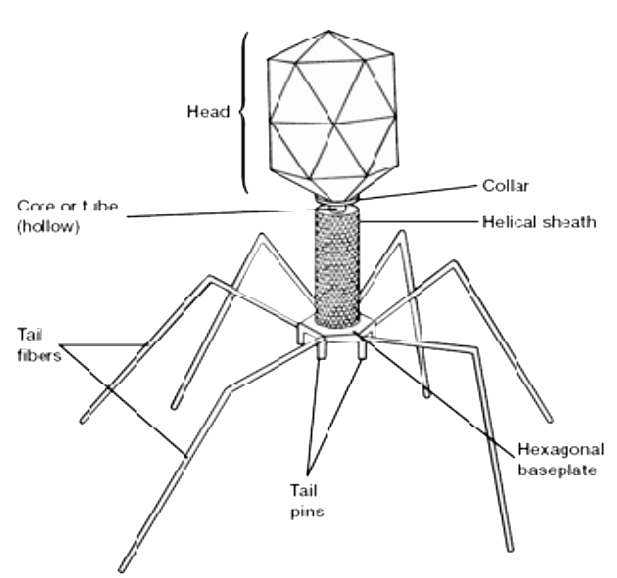
Virus particle structure
T4 is a relatively large phage, at approximately 90 nm wide and 200 nm long (most phages range from 25 to 200 nm in length). The DNA genome is held in an icosahedral head, also known as a capsid. The T4’s tail is hollow so that it can pass its nucleic acid into the cell it is infecting after attachment. The tail attaches to a host cell with the help of tail fibres. The tail fibres are also important in recognizing host cell surface receptors, so they determine if a bacterium is within the phage's host range.
The structure of the 6 megadalton T4 baseplate that comprises 127 polypeptide chains of 13 different proteins (gene products 5, 5.4, 6, 7, 8, 9, 10, 11, 12, 25, 27, 48 and 53) has recently been described in atomic detail. An atomic model of the proximal region of the tail tube formed by gp54 and the main tube protein gp19 have also been created. The tape measure protein gp29 is present in the baseplate-tail tube complexes, but it could not be modeled.
Infection process
The T4 phage initiates an E. coli infection by binding OmpC porin proteins and Lipopolysaccharide (LPS) on the surface of E. coli cells with its long tail fibers (LTF). A recognition signal is sent through the LTFs to the baseplate. This unravels the short tail fibers (STF) that bind irreversibly to the E. coli cell surface. The baseplate changes conformation and the tail sheath contracts, causing GP5 at the end of the tail tube to puncture the outer membrane of the cell. The lysozyme domain of GP5 is activated and degrades the periplasmic peptidoglycan layer. The remaining part of the membrane is degraded and then DNA from the head of the phage can travel through the tail tube and enter the E. coli cell.
Reproduction
The lytic lifecycle (from entering a bacterium to its destruction) takes approximately 30 minutes (at 37 °C) and consists of:
After the life cycle is complete, the host cell bursts open and ejects the newly built viruses into the environment, destroying the host cell. T4 has a burst size of approximately 100-150 viral particles per infected host. Complementation, deletion, and recombination tests can be used to map out the rII gene locus by using T4. These bacteriophage infect a host cell with their information and then blow up the host cell, thereby propagating themselves.
Replication and packaging
The rate of DNA replication in a living cell was measured as the rate of phage T4 DNA elongation in phage-infected E. coli. During the period of exponential DNA increase at 37 °C, the rate was 749 nucleotides per second. The mutation rate per base pair per replication during phage T4 DNA synthesis is 1.7 per 10−8, a highly accurate DNA copying mechanism, with only 1 error in 300 copies. The phage also codes for unique DNA repair mechanisms. The T4 DNA packaging motor has been found to load DNA into phage capsids at a rate up to 2000 base pairs per second. The power involved, if scaled up in size, would be equivalent to that of an average automobile engine.
Multiplicity reactivation
Multiplicity reactivation (MR) is the process by which two or more virus genomes, each containing inactivating genome damage, can interact within an infected cell to form a viable virus genome. Salvador Luria, while studying UV irradiated phage T4 in 1946, discovered MR and proposed that the observed reactivation of damaged phage occurs by a recombination mechanism.(see refs.) This preceded the confirmation of DNA as the genetic material in 1952 in related phage T2 by the Hershey–Chase experiment.
As remembered by Luria (1984, pg. 97) the discovery of reactivation of irradiated phage (referred to as "multiplicity reactivation") immediately started a flurry of activity in the study of repair of radiation damage within the early phage group (reviewed by Bernstein in 1981). It turned out later that the repair of damaged phage by mutual help that Luria had discovered was only one special case of DNA repair. Cells of all types, not just, bacteria and their viruses, but all organisms studied, including humans, are now known to have complex biochemical processes for repairing DNA damages (see DNA repair). DNA repair processes are also now recognized as playing critical roles in protecting against aging, cancer, and infertility.
MR is usually represented by "survival curves" where survival of plaque forming ability of multiply infected cells (multicomplexes) is plotted against dose of genome damaging agent. For comparison, the survival of phage plaque forming ability of singly infected cells (monocomplexes) is also plotted against dose of genome damaging agent. The top figure shows the survival curves for phage T4 multicomplexes and monocomplexes with increasing dose of UV light. Since survival is plotted on a log scale it is clear that survival of multicomplexes exceeds that of monocomplexes by very large factors (depending on dose). The UV inactivation curve for multicomplexes has an initial shoulder. Other phage T4 DNA damaging agents with shoulders in their multicomplex survival curves are X-rays and ethyl methane sulfonate (EMS). The presence of a shoulder has been interpreted to mean that two recombinational processes are used. The first one repairs DNA with high efficiency (in the "shoulder"), but is saturated in its ability as damage increases; the second pathway functions at all levels of damage. Surviving T4 phage released from multicomplexes show no increase in mutation, indicating that MR of UV irradiated phage is an accurate process.
The bottom figure shows the survival curves for inactivation of phage T4 by the DNA damaging agent mitomycin C (MMC). In this case the survival curve for multicomplexes has no initial shoulder, suggesting that only the second recombinational repair process described above is active. The efficiency of repair by this process is indicated by the observation that a dose of MMC that allows survival of only 1 in 1,000 monocomplexes allows survival of about 70% of multicomplexes. Similar multicomplex survival curves (without shoulders) were also obtained for the DNA damaging agents P32 decay, psoralen plus near-UV irradiation (PUVA), N-methyl-N'-nitro-N-nitrosoguanidine (MNNG), methyl methane sulfonate (MMS) and nitrous acid.
Several of the genes found to be necessary for MR in phage T4 proved to be orthologs for genes essential for recombination in prokaryotes, eukaryotes and archaea. This includes, for instance, T4 gene uvsX which specifies a protein that has three-dimensional structural homology to RecA from Escherichia coli and the homologous protein RAD51 in eukaryotes and RadA in archaea. It has been suggested that the efficient and accurate recombinational repair of DNA damages during MR may be analogous to the recombinational repair process that occurs during meiosis in eukaryotes.
History
The specific time and place of T4 phage isolation remains unclear, though they were likely found in sewage or fecal material. T4 and similar phages were described in a paper by Thomas F. Anderson, Max Delbrück, and Milislav Demerec in November 1944.
A number of Nobel Prize winners worked with phage T4 or T4-like phages including Max Delbrück, Salvador Luria, Alfred Hershey, James D. Watson, and Francis Crick. Other important scientists who worked with phage T4 include Michael Rossmann, Seymour Benzer, Bruce Alberts, Gisela Mosig, Richard Lenski, and James Bull.
