 | ||
A transmembrane protein (TP) is a type of integral membrane protein that spans the entirety of the biological membrane to which it is permanently attached. Many transmembrane proteins function as gateways to permit the transport of specific substances across the biological membrane. They frequently undergo significant conformational changes to move a substance through the membrane.
Contents
- Classification by structure
- Classification by topology
- 3D structure
- Stability of helical transmembrane proteins
- Folding of helical transmembrane proteins
- Stability and folding of barrel transmembrane proteins
- Light absorption driven transporters
- Oxidoreduction driven transporters
- Electrochemical potential driven transporters
- P P bond hydrolysis driven transporters
- Porters uniporters symporters antiporters
- Alpha helical channels including ion channels
- Enzymes
- Proteins with alpha helical transmembrane anchors
- barrels composed of a single polypeptide chain
- barrels composed of several polypeptide chains
- References
Transmembrane proteins are polytopic proteins that aggregate and precipitate in water. They require detergents or nonpolar solvents for extraction, although some of them (beta-barrels) can be also extracted using denaturing agents.
The other type of integral membrane protein is the integral monotopic protein that is also permanently attached to the cell membrane but does not pass through it.
Classification by structure
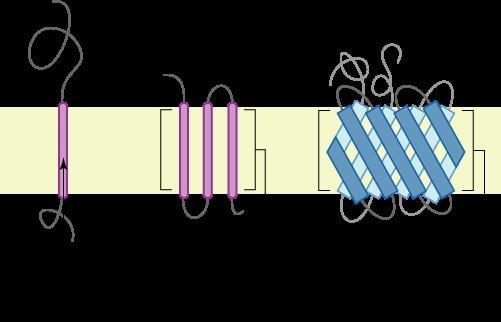
There are two basic types of transmembrane proteins: alpha-helical and beta-barrels. Alpha-helical proteins are present in the inner membranes of bacterial cells or the plasma membrane of eukaryotes, and sometimes in the outer membranes. This is the major category of transmembrane proteins. In humans, 27% of all proteins have been estimated to be alpha-helical membrane proteins. Beta-barrel proteins are so far found only in outer membranes of gram-negative bacteria, cell wall of gram-positive bacteria, and outer membranes of mitochondria and chloroplasts. All beta-barrel transmembrane proteins have simplest up-and-down topology, which may reflect their common evolutionary origin and similar folding mechanism.
Classification by topology
This classification refers to the position of the N- and C-terminal domains. Types I, II, and III are single-pass molecules, while type IV are multiple-pass molecules. Type I transmembrane proteins are anchored to the lipid membrane with a stop-transfer anchor sequence and have their N-terminal domains targeted to the ER lumen during synthesis (and the extracellular space, if mature forms are located on plasmalemma). Type II and III are anchored with a signal-anchor sequence, with type II being targeted to the ER lumen with its C-terminal domain, while type III have their N-terminal domains targeted to the ER lumen. Type IV is subdivided into IV-A, with their N-terminal domains targeted to the cytosol and IV-B, with an N-terminal domain targeted to the lumen. The implications for the division in the four types are especially manifest at the time of translocation and ER-bound translation, when the protein has to be passed through the ER membrane in a direction dependent on the type.
3D structure
Membrane protein structures can be determined by X-ray crystallography, electron microscopy or NMR spectroscopy. The most common tertiary structures of these proteins are transmembrane helix bundle and beta barrel. The portion of the membrane proteins that are attached to the lipid bilayer (see annular lipid shell) consist mostly of hydrophobic amino acids.
Membrane proteins have hydrophobic surfaces, are relatively flexible and are expressed at relatively low levels. This creates difficulties in obtaining enough protein and then growing crystals. Hence, despite the significant functional importance of membrane proteins, determining atomic resolution structures for these proteins is more difficult than globular proteins. As of January 2013 less than 0.1% of protein structures determined were membrane proteins despite being 20-30% of the total proteome. Due to this difficulty and the importance of this class of proteins methods of protein structure prediction based on hydropathy plots, the positive inside rule and other methods have been developed.
Stability of α-helical transmembrane proteins
Transmembrane α-helical proteins are unusually stable judging from thermal denaturation studies, because they do not unfold completely within the membranes (the complete unfolding would require breaking down too many α-helical H-bonds in the nonpolar media). On the other hand, these proteins easily misfold, due to non-native aggregation in membranes, transition to the molten globule states, formation of non-native disulfide bonds, or unfolding of peripheral regions and nonregular loops that are locally less stable.
It is also important to properly define the unfolded state. The unfolded state of membrane proteins in detergent micelles is different from that in the thermal denaturation experiments. This state represents a combination of folded hydrophobic α-helices and partially unfolded segments covered by the detergent. For example, the "unfolded" bacteriorhodopsin in SDS micelles has four transmembrane α-helices folded, while the rest of the protein is situated at the micelle-water interface and can adopt different types of non-native amphiphilic structures. Free energy differences between such detergent-denatured and native states are similar to stabilities of water-soluble proteins (< 10 kcal/mol).
Folding of α-helical transmembrane proteins
Refolding of α-helical transmembrane proteins in vitro is technically difficult. There are relatively few examples of the successful refolding experiments, as for bacteriorhodopsin. In vivo, all such proteins are normally folded co-translationally within the large transmembrane translocon. The translocon channel provides a highly heterogeneous environment for the nascent transmembane α-helices. A relatively polar amphiphilic α-helix can adopt a transmembrane orientation in the translocon (although it would be at the membrane surface or unfolded in vitro), because its polar residues can face the central water-filled channel of the translocon. Such mechanism is necessary for incorporation of polar α-helices into structures of transmembrane proteins. The amphiphilic helices remain attached to the translocon until the protein is completely synthesized and folded. If the protein remains unfolded and attached to the translocon for too long, it is degraded by specific "quality control" cellular systems.
Stability and folding of β-barrel transmembrane proteins
Stability of β-barrel transmembrane proteins is similar to stability of water-soluble proteins, based on chemical denaturation studies. Their folding in vivo is facilitated by water-soluble chaperones, such as protein Skp [1].
Light absorption-driven transporters
Oxidoreduction-driven transporters
Electrochemical potential-driven transporters
P-P-bond hydrolysis-driven transporters
Porters (uniporters, symporters, antiporters)
Alpha-helical channels including ion channels
Enzymes
Proteins with alpha-helical transmembrane anchors
β-barrels composed of a single polypeptide chain
Note: n and S are, respectively, the number of beta-strands and the "shear number" of the beta-barrel
β-barrels composed of several polypeptide chains
See also Gramicidin A [56], a peptide that forms a dimeric transmembrane β-helix. It is also secreted by Gram-positive bacteria.
