Possible place of origin Horn of Africa Descendants E-M2, E-M329 | Ancestor E-P2 Defining mutations L222.1, V38, V100 | |
 | ||
Possible time of origin approx 25,000-30,000 years BP | ||
Haplogroup E-V38 is a human Y-chromosome DNA haplogroup. It is primarily distributed in Africa. E-V38 has two basal branches, E-M329 (formerly E1b1c) and E-M2 (formerly E1b1a); the E-M329 subclade is today almost exclusively found in Ethiopia. E-M2 is the predominant subclade in Western Africa, Central Africa, Southern Africa and the African Great Lakes, and occurs at low frequencies in other areas. E-M2 has several subclades, but many members are included in either E-L485 or E-U175.
Contents
Origins
The discovery of two SNPs (V38 and V100) by Trombetta et al. (2011) significantly redefined the E-V38 phylogenetic tree. This led the authors to suggest that E-V38 may have originated in East Africa. V38 joins the West African-affiliated E-M2 and the northern East African-affiliated E-M329 with an earlier common ancestor who, like E-P2, may have also originated in East Africa. It is possible that soon after the evolution of E-V38, trans-Saharan migrants carried the E-V38 marker to North and Central Africa and/or West Africa where the more common E-M2 marker later arose and became prolific within the last 20,000-30,000 years.
The downstreams SNP E-M180 possibly originated on the moist south-central Saharan savannah/grassland of northern West Africa during the early Holocene period. Much of the population that carried E-M2 retreated to southern West Africa with the drying of the Sahara. These later people migrated from Southeastern Nigeria and Cameroon ~8.0 kya to Central Africa, East Africa, and Southern Africa causing or following the Bantu expansion. According to Wood et al. (2005) and Rosa et al. (2007), such population movements from West Africa changed the pre-existing population Y chromosomal diversity in Central, Southern and southern East Africa, replacing the previous haplogroups frequencies in these areas with the now dominant E1b1a1 lineages. Traces of earlier inhabitants, however, can be observed today in these regions via the presence of the Y DNA haplogroups A1a, A1b, A2, A3, and B-M60 that are common in certain populations, such as the Mbuti and Khoisan.
Distribution
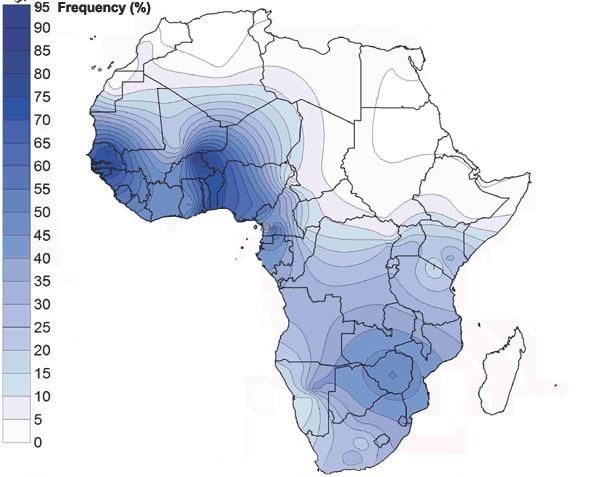
This haplogroup's frequency and diversity are highest in the West Africa region. Within Africa, E-V38 displays a west-to-east as well as a south-to-north clinal distribution. In other words, the frequency of the haplogroup decreases as one moves from western and southern Africa toward the eastern and northern parts of the continent.
Populations on the northern fringes of West Africa, central Eastern Africa and Madagascar have tested at more moderate frequencies.
E-V38 is found at low to moderate frequencies in North Africa, and northern East Africa. The some of the lineages found in these areas are possibly due to the Bantu expansion or other migrations. The E-M2 marker that appeared in North African samples may stem from recent acquisitions. However, the discovery in 2011 of the E-V38 marker that predates E-M2 has led Trombetta et al. to suggest that E-V38 may have originated in East Africa (please refer to the Origins section for the details).
Outside of Africa, E-V38 has been found at low frequencies. The clade has been found at low frequencies in West Asia. A few isolated occurrences of E-V38 have also been observed among populations in Southern Europe, such as Croatia, Malta, Spain and Portugal.
The Trans-Atlantic slave trade brought people to North America, Central America and South America including the Caribbean. Consequently, the haplogroup is often observed in the United States populations in men who self-identify as African Americans. It has also been observed in a number of populations in Mexico, the Caribbean, Central America, and South America among people of African descent.
E1b1a1
E1b1a1 is defined by markers DYS271/M2/SY81, M291, P1/PN1, P189, P293, V43, and V95.
E1b1a1a1
E1b1a1a1 is commonly defined by M180/P88. The basal subclade is quite regularly observed in V38+ samples.
E1b1a1a1a
E1b1a1a1a is defined by marker M58. 5% (2/37) of the town Singa-Rimaïbé, Burkina Faso tested positive for E-M58. 15% (10/69) of Hutus in Rwanda tested positive for M58. Three South Africans tested positive for this marker. One Carioca from Rio de Janeiro, Brazil tested positive for the M58 SNP. The place of origin and age is unreported.
E1b1a1a1b
E1b1a1a1b is defined by M116.2, a private marker. A single carrier was found in Mali.
E1b1a1a1c
E1b1a1a1c is defined by private marker M149. This marker was found in a single South African.
E1b1a1a1d
E1b1a1a1d is defined by a private marker M155. It is known from a single carrier in Mali.
E1b1a1a1e
E1b1a1a1e is defined by markers M10, M66, M156 and M195. Wairak people in Tanzania tested 4.6% (2/43) positive for E-M10. E-M10 was found in a single person of the Lissongo group in the Central African Republic and two members in a "Mixed" population from the Adamawa region.
E1b1a1a1f
E1b1a1a1f is defined by L485. The basal node E-L485* appears to be somewhat uncommon but has not been sufficiently tested in large populations. The ancestral L485 SNP (along with several of its subclades) was very recently discovered. Some of these SNPs have little or no published population data and/or have yet to receive nomenclature recognition by the YCC.
E1b1a1a1g
E1b1a1a1g (YCC E1b1a8) is defined by marker U175. The basal E-U175* is extremely rare. Montano et al. (2011) only found one out of 505 tested African subjects who was U175 positive but negative for U209. Brucato et al. found similarly low frequencies of basal E-U175* in subjects in the Ivory Coast and Benin. Veeramah et al. (2010) found U175 in tested Annang (45.3%), Ibibio (37%), Efik (33.3%), and Igbo (25.3%) but did not test for U209.
The supposed "Bantu haplotype" found in E-U175 carriers is "present at appreciable frequencies in other Niger–Congo languages speaking peoples as far west as Guinea-Bissau". This is the modal haplotype of STR markers that is common in carriers of E-U175.
E1b1a1a1g has several subclades.
E1b1a1a1h
E1b1a1a1h is defined by markers P268 and P269. It was first reported in a person from the Gambia.
E1b1a2
E1b1a2 is defined by the SNP mutation M329. The majority of the few cases so far observed have been found in East Africa. Semino et al. (2004) found 2 Ethiopian Oromo in a study of >2400 individuals, including 78 Oromo. Cadenas et al. (2007) found 1 case in Qatar, out of 72 people tested there in that study.
Phylogenetic history
Prior to 2002, there were in academic literature at least seven naming systems for the Y-Chromosome Phylogenetic tree. This led to considerable confusion. In 2002, the major research groups came together and formed the Y-Chromosome Consortium (YCC). They published a joint paper that created a single new tree that all agreed to use. Later, a group of citizen scientists with an interest in population genetics and genetic genealogy formed a working group to create an amateur tree aiming at being above all timely. The table below brings together all of these works at the point of the landmark 2002 YCC Tree. This allows a researcher reviewing older published literature to quickly move between nomenclatures.
Research publications
The following research teams per their publications were represented in the creation of the YCC tree.
Tree
This phylogenetic tree of haplogroup subclades is based on the Y-Chromosome Consortium (YCC) 2008 Tree, the ISOGG Y-DNA Haplogroup E Tree, and subsequent published research.
