Stable release 4.14.4 / 2008 License Proprietary | ||
 | ||
Developer(s) Daniel Huson and David Bryant Operating system Website | ||
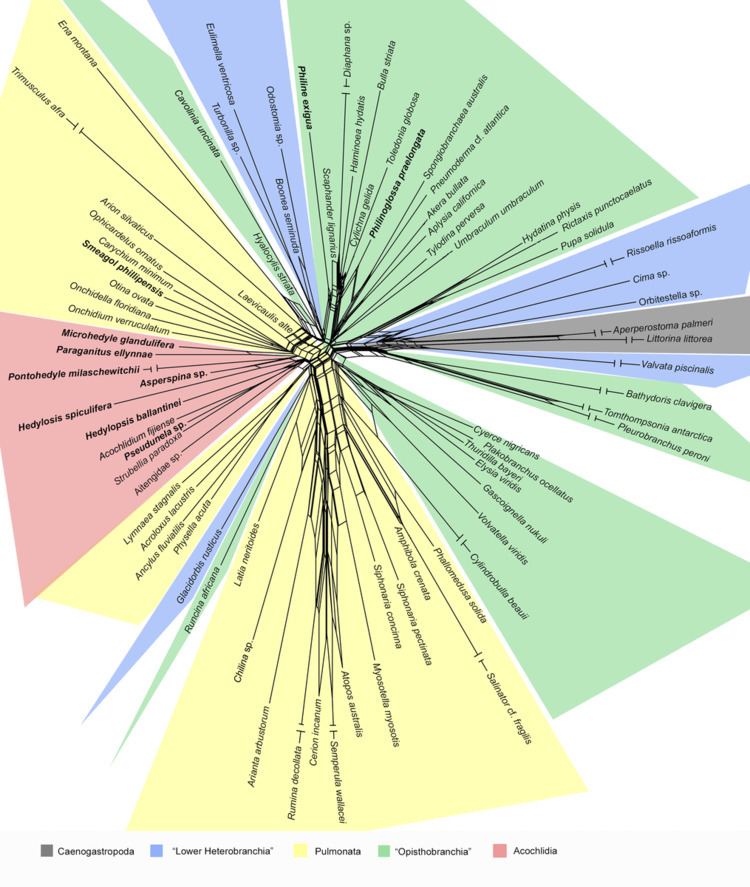
SplitsTree is a popular program for inferring phylogenetic trees or, more generally, phylogenetic networks from various types of data such as a sequence alignment, a distance matrix or a set of trees. SplitsTree implements published methods such as split decomposition neighbor-net, consensus networks, super networks methods or methods for computing hybridization or simple recombination networks.
References
SplitsTree Wikipedia(Text) CC BY-SA
