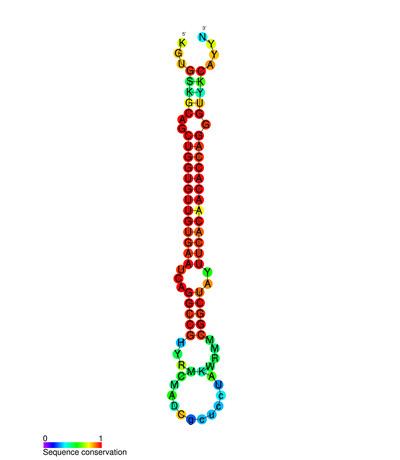
miR-138 is a family of microRNA precursors found in animals, including humans. MicroRNAs are typically transcribed as ~70 nucleotide precursors and subsequently processed by the Dicer enzyme to give a ~22 nucleotide product. The excised region or, mature product, of the miR-138 precursor is the microRNA mir-138.
miR-138 has been used as an example of the post-transcriptional regulation of miRNA, due to the finding that while the precursor is expressed ubiquitously, the mature product is found only in specific cell types.
The presence of miR-138 has been detected experimentally in humans (Homo sapiens) and in different animals including house mouse (Mus musculus), brown rat (Rattus norvegicus), platypus (Ornithorhynchus anatinus), Carolina anole(Anolis carolinensis), cattle (Bos taurus), common carp (Cyprinus carpio), dog (Canis familiaris), Chinese hamster (Cricetulus griseus), zebrafish (Danio rerio), red junglefowl (Gallus gallus), western gorilla (Gorilla gorilla), gray short-tailed opossum (Monodelphis domestica), Oryzias latipes, sea lamprey (Petromyzon marinus), Tasmanian devil (Sarcophilus harrisii), wild boar (Sus scrofa) and zebra finch (Taeniopygia guttata).
It is also predicted computationally that the miR-138 gene exists in the genome of other animals including horse (Equus caballus), rhesus macaque (Macaca mulatta), takifugu rubripes (Fugu rubripes), Bornean orangutan (Pongo pygmaeus), common chimpanzee (Pan troglodytes), Tetraodon nigroviridis and western clawed frog (Xenopus tropicalis).
In human genome, there are two miR-138 associated genes and they are not located in any cluster. More precisely, the miR-138-1 gene is in region 5 at 3p21.3 and miR-138-2 is located on chromosome 16 (16q13).
In adult mice, miR-138 is only expressed in brain tissue. Its expression is not uniform throughout the brain but restricted to distinct neuronal populations. On the contrary, its precursor, pre-miR-138-2, is ubiquitously expressed throughout all tissues, which suggests that the expression of miRNAs can be regulated at the post-transcription level.
In the zebrafish, miR-138 is expressed in specific domains in the heart and is required to establish appropriate chamber-specific gene expression patterns.
Targets and function
Since the identification of miR-138, a number of targets have been found and some of them have been verified experimentally. It has been proven that miR-138 is involved in different pathways. Furthermore, it is in relation with various types of cancer.
HIF-1aHypoxia-inducible factor-1alpha (HIF-1a), one of the key regulators in cancer cells, has been shown to be one target of miR-138.
VIM, ZEB2, EZH2 and head and neck cancersDownregulation of miR-138 has been reported in several types of cancers, including HNSCC(head and neck squamous cell carcinoma). It is suggested that miR-138 is a multi-functional molecular regulator and plays major roles in EMT (epithelial-mesenchymal transition) and in HNSCC progression. A number of miR-138 target genes have been identified to be associated with EMT, including VIM (
vimentin), ZEB2 (zinc finger E-box-binding homeobox 2) and EZH2 (enhancer of zeste homologue 2).
CCND1 and nasopharyngeal carcinomamiR-138 is commonly underexpressed in nasopharyngeal carcinoma (NPC) specimens and NPC cell lines.
Cyclin D1 (CCND1), which is widely upregulated in NPC tumors, is found as a direct target of miR-138. Therefore, miR-138 might be a tumor suppressor in NPC, which is exerted partially by inhibiting CCND1 expression.
BCR-ABL and CCND3BCR (breakpoint cluster region)-ABL (c-abl oncogene 1, non-receptor tyrosine kinase)/
GATA1/miR-138 mini circuitry contributes to the leukemogenesis of chronic myeloid leukemia (CML). ABL and BCR-ABL are the target genes of miR-138, which binds to the
coding region instead of
three prime untranslated region (3'UTR). miR-138 can negatively regulate another gene CCND3 via binding to its 3'-UTR. The expression of miR-138 is activated by GATA1, which in turn is repressed by BCR-ABL. Therefore, miR-138, by virtue of a BCR-ABL/GATA1/miR-138 circuitry, is a tumor suppressor miRNA implicated in the pathogenesis of CML and its clinical response to
imatinib.
H2AX and DNA damage repairmir-138 is linked with DNA damage repair. It can directly target the
histone H2AX 3'UTR, reduce histone H2AX expression and induce chromosomal instability after DNA damage.
ALDH1A2 and CSPG2In zebrafish, the mature form of miR-138 regulates gene expression influencing cardiac development. miR-138 helps establish discrete domains of gene expression during cardiac morphogenesis by targeting multiple members of a common pathway. It has been experimentally verified that miR-138 can negatively regulate
aldh1a2, encoding
retinoic acid (RA) dehydrogenase (Raldh2), by targeting the binding site in the 3'UTR of its mRNA. Another putative target of miR-138 is
cspg2.
Regulation of sleepIn rats, miR-138, let-7b, and miR-125a are expressed at different times and in different structures in the brain and likely play a role in the regulation of sleep.
Brain cancermiR-138 has been found to be significantly linked with the formation and growth of
Gliomas, from Cancerous Stem Cells (CSC). In vitro inhibition of miR-138 prevents tumour sphere formation. Furthermore, its high expression in Glioma makes it a potential biomarker for CSC.
Rhoc, ROCK2 and Tongue cancerTumour metastasis concerning the Tongue Squamous Cell Carcinoma (TSCC) can be regulated via the expression of 2 key genes in Rho GTPase signaling pathway :
RhoC and ROCK2 (
Rho-associated protein kinase 2). Thus, by targeting the 3' untranslated region of those genes, mir-138 is able to reduce their expression and by this mean, to destroy TSCC ability migrate and invade.