Entrez 4350 | Ensembl ENSG00000103152 | |
 | ||
Aliases MPG, AAG, ADPG, APNG, CRA36.1, MDG, Mid1, PIG11, PIG16, anpg, N-methylpurine DNA glycosylase External IDs MGI: 97073 HomoloGene: 1824 GeneCards: MPG | ||
DNA-3-methyladenine glycosylase also known as 3-alkyladenine DNA glycosylase (AAG) or N-methylpurine DNA glycosylase (MPG) is an enzyme that in humans is encoded by the MPG gene.
Contents
- Function
- Alkylation repairing activity
- ODG activity
- Structure
- Isoform 2
- Isoform 3
- Isoform 4
- Substrate recognition
- Nucleotide flipping and fixation
- Nucleotide release
- Location
- Clinical significance
- Model organisms
- References
Alkyladenine DNA glycosylase is a specific type of DNA glycosylase. This subfamily of monofunctional glycosylases is involved in the recognition of a variety of base lesions, including alkylated and deaminated purines, and initiating their repair via the base excision repair pathway. To date, the human AAG (hAAG) is the only glycosylase identified that excises alkylation-damaged purine bases.
Function
DNA bases are subject to a large number of anomalies: spontaneous alkylation or oxidative deamination. It is estimated that 104 mutations appear in a typical human cell per day. Albeit it seems to be an insignificant amount considering the extension of the DNA (1010 nucleotides), these mutations lead to changes in the structure and coding potential of the DNA, affecting processes of replication and transcription.
3-Methyladenine DNA glycosilases are able to initiate the base excision repair (BER) of a wide range of substrate bases that, due to their chemical reactivity, suffer inevitable modifications resulting in different biological outcomes. DNA repair mechanisms take on a vital role in maintaining the genomic integrity of cells from different organisms, in particular 3-Methyladenine DNA glycosylases are found in bacteria, yeast, plants, rodents and humans. Therefore, there are different subfamilies of this enzyme, such as the Human Alkyladenine DNA Glycosylase (hAAG), that act on other damaged DNA bases apart from 3-MeA
Alkylation repairing activity
In cells, AAG is the enzyme responsible for recognition and initiation of the repair, via catalysing the hydrolysis of the N-glycosidic bond to release the alkylation-damaged purine bases. Specifically, hAAG is able to efficiently identify and excise 3-methyladenine, 7-methyladenine, 7-methylguanine, 1N-ethenoadenine and hypoxanthine.
ODG activity
Oxanine DNA Glycolase (ODG) activity is the capability of some DNA Glycosylases of repairing Oxanines (Oxa), a deamined base lesion in which the N1-nitrogen is replaced by oxygen. Among the known human DNA glycosylases tested, the human alkyladenine DNA glycosylase (AAG) also shows ODG activity.
Contrary to the alkylation repairing activity, which is only able to act against purine bases, the hAAG is able to excise Oxa from all of four Oxa-containing double stranded base pairs, Cyt/Oxa, Thy/Oxa, Ade/Oxa, and Gua/Oxa, showing no particular preference by any of the bases. In addition hAAG is capable of removing Oxa from single-stranded Oxa- containing DNA. This occurs because the ODG activity of the hAAG does not require a complementary strand.
Structure
Alkyladenine DNA glycosylase is a monomeric protein compounded by 298 amino acids, with a formula weight of 33kDa. Its canonical primary structure consists of the following sequence. However, also other functional isoforms have been found.
Isoform 2
The sequence of this isoform differs from the canonical sequence as follows:
Aminoacids 1-12: MVTPALQMKKPK → MPARSGA
Aminoacids 195-196: QL →HV
Isoform 3
The sequence of this isoform differs from the canonical sequence in a similar way as the isoform 2:
Aminoacids 1-12: MVTPALQMKKPK → MPARSGA
Isoform 4
The sequence of this isoform misses the aminoacids 1-17.
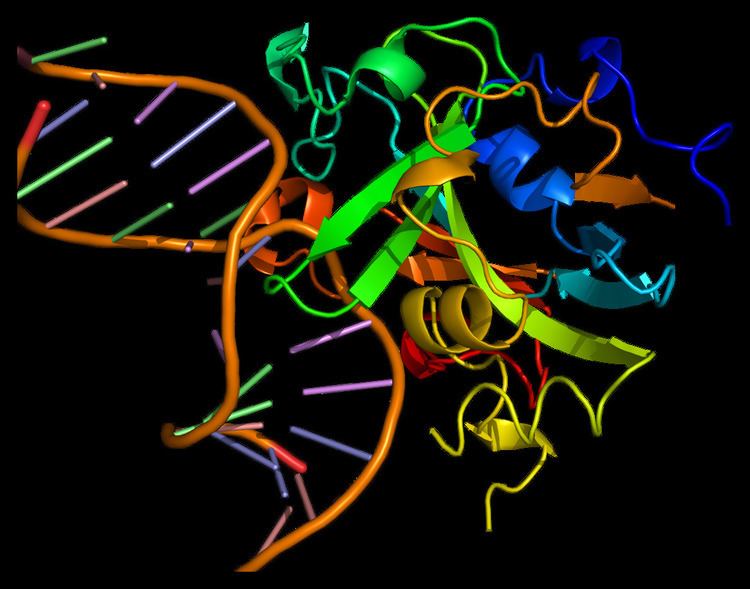
It folds into a single domain of mixed α/β structure with seven α helices and eight β strands. The core of the protein consists of a curved, antiparallel β sheet with a protruding β hairpin (β3β4) that inserts into the minor groove of the bound DNA. A series of α helices and connecting loops form the remainder of the DNA binding interface. It lacks the helix-hairpin-helix motif associated with other base excision-repair proteins and, in fact, it does not resemble any other model in the Protein Data Bank.
Substrate recognition
Alkyladenine DNA glycosylase is part of the family of enzymes that follow the BER, acting on specific substrates according to BER steps.
The process of recognition of damaged bases involves initial non-specific binding followed by diffusion along the DNA. Formed the AAG-DNA complex, a redundant process of search occurs because of the long lifetime of this complex, while hAAG search many adjacent sites in a DNA molecule in a single binding. This provides ample opportunity to recognize and excise lesions that minimally perturb the structure of the DNA.
Due to its broad specificity, the hAAG performs the substrate selection through a combination of selectivity filters.
Nucleotide flipping and fixation
Its structure contains an antiparallel β sheet with protruding β hairpin (β3β4) that inserts into the minor groove of the bound DNA. This group is unique for the human cells and displaces the selected nucleotide targeted for base excision by flipping it. The nucleotide is secured into the enzyme binding pocket where the active site is found, and is fixed by the amino acids Arg182, Glu125 and Ser262. Also other bonds are formed to bordering nucleotides to stabilize the structure.
The groove in the double helix of DNA left by the flipped-out abasic nucleotide is filled with the lateral chain of the amino acid Tyr162, making no specific contacts with the surrounding bases.
Nucleotide release
Activated by nearby aminoacids, a water molecule attacks the N-Glycosydic bound releasing the alkylated base via a backside displacement mechanism.
Location
Human alkyladenine DNA glycosylase localizes to the mitochondria, nucleus and cytoplasm of human cells. Despite only being found in human cells, some functionally equivalent enzymes have been found in other species, but with significantly different structures, such as E. coli DNA-3-methyladenine glycosylase.
Clinical significance
According to the mechanism used by Human Alkyladenine DNA Glycosylase, a defect in the DNA repair pathways leads to cancer predisposition. HAAG follows the BER steps so that means that an incorrect role of BER genes could contribute to the development of cancer. Concretely, a bad activity of hAAG may be associated with cancer risk in BRCA1 and BRCA2 mutation carriers.
Model organisms
Model organisms have been used in the study of MPG function. A conditional knockout mouse line called Mpgtm1a(EUCOMM)Wtsi was generated at the Wellcome Trust Sanger Institute. Male and female animals underwent a standardized phenotypic screen to determine the effects of deletion. Additional screens performed: - In-depth immunological phenotyping
