 | ||
A transcriptome in vivo analysis tag (TIVA tag) is a multifunctional, photoactivatable mRNA-capture molecule designed for isolating mRNA from a single cell in complex tissues.
Contents
Background
A transcript is an RNA molecule that is copied or transcribed from a DNA template. A transcript can be further processed by alternative splicing, which is the retention of different combinations of exons. These unique combinations of exons are termed RNA transcript isoforms. The transcriptome is a set of all RNA, including rRNA, mRNA, tRNA, and non-coding RNA. Specifically mRNA transcripts can be used to investigate differences in gene expression patterns. Transcriptome profiling is determining the composition of transcripts and their relative expression levels in a given reference set of cells. This analysis involves characterization of all functional genomic elements, coding and non-coding.
The current RNA capture methods involve sorting cells in suspension from acutely dissociated tissue, and thus can lose information about cell morphology and microenvironment. Transcript abundance and isoforms are significantly different across tissues and are continually changing throughout an individual’s life. Gene expression is highly tissue specific, therefore with traditional RNA capture methods one must be cautious in the interpretation of gene expression patterns, as they often reflect expression of a heterogeneous mix of cell populations. Even in the same cell type, tissue measurements, where a population of cells is obtained, mask both low-level mRNA expression in single cells and variation in expression between cells. The photoactivatable TIVA tag is engineered to capture the mRNA of a single cell in complex tissues.
Chemical Structure
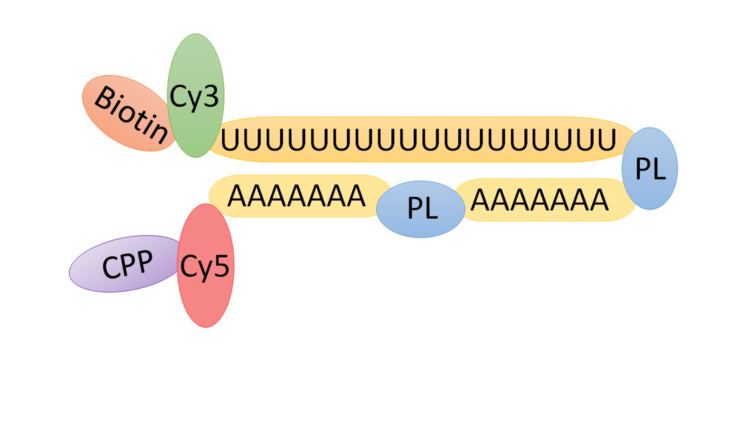
TIVA tags are created initially via solid-phase synthesis with the cell-penetrating peptide conjugated afterwards. The functional components of the tag can be summarized as following:
Tissue preparation
Tissue fixation is performed by chemical fixation using formalin. This prevents the postmortem degeneration of the tissue and hardens soft tissue. The tissue is dehydrated using ethanol and the alcohol is cleared using an organic solvent such as xylene. The tissue is embedded in parafin which infiltrates the microscopic spaces present throughout the tissue. The embedded tissue is sliced using a microtome and subsequently stained to produce contrast needed to visualize the tissue.
Loading of the TIVA tag into cells and validation
A cell saline buffer containing the TIVA tag is added to the coverslip and incubated. During the incubation period, the TIVA tag penetrates the cell membrane via the CPP that is bound to it. Subsequently, the cytosolic environment cleaves the CPP and the TIVA tag is trapped inside the cell. After incubation, the coverslip is rinsed twice with cell saline buffer and then transferred to an imaging chamber. Using a confocal microscope, loading of the tag is confirmed by detecting the Cy5 signal at a wavelength of 561 nm.
Photoactivation of the TIVA tag in target cell and validation
Photolysis is performed resulting in photoactivation of the TIVA tag in the target cell or cells. Specifically, uncaging of the TIVA tag is accomplished using a 405-nm laser while measuring FRET excited by 514 nm. During this process, the mRNA-capturing moiety is released and subsequently anneals to the poly(A) tail of cellular mRNA. To confirm that the cell is not damaged during photolysis, the cell is imaged with the confocal microscope.
Extraction, lysis of target cell and affinity purification of TIVA tag
Using a glass pipette, the photolysed cell is isolated by aspiration. Cells are lysed and affinity purification is performed using streptavidin-coated beads that bind, immobilize and purify the biotinylated TIVA tag.
RNA-seq analysis
RNA-seq uses reverse transcriptase to convert the mRNA template to cDNA. During library preparation, the cDNA is fragmented into small pieces, which then serve as the template for sequencing. After sequencing RNA-seq analysis can then be performed.
