Symbol SI HUGO 10856 PDB 3LPO | Entrez 6476 OMIM 609845 RefSeq NM_001041 | |
 | ||
Sucrase-isomaltase (EC 3.2.1.10, is a glucosidase enzyme located in on the brush border of the small intestine. Sucrase-isomaltase is a type II transmembrane glycoprotein located in the brush border of the small intestine. It has preferential expression in the apical membranes of enterocytes. The enzyme’s purpose is to digest dietary carbohydrates such as starch, glucose, and isomaltose. By further processing the broken-down products, energy in the form of ATP can be generated.
Contents
Nomenclature
The systematic name of systematic name of sucrase-isomaltase is oligosaccharide 6-alpha-glucohydrolase. This enzyme is also known as:
Mechanism
This enzyme catalyses the following chemical reaction
Hydrolysis of (1->6)-alpha-D-glucosidic linkages in some oligosaccharides produced from starch and glycogen by enzyme EC 3.2.1.1.Hydrolysis uses water to cleave chemical bonds. Sucrase-isomaltase’s mechanism results in a net retention of configuration at the anomeric center.
Structure
Sucrase-isomaltase consists of two enzymatic subunits: sucrase and isomaltase. The subunits originate from a polypeptide precursor, pro-SI. By heterodimerizing the two subunits, the sucrase-isomaltase complex is formed. The enzyme is anchored in the intestinal brush border membrane by a hydrophobic segment located near the N-terminal of the isomaltase subunit. Before the enzyme is anchored to the membrane, pro-SI is mannose-rich and glycosylated; it moves from the ER to the Golgi, where it becomes a protein complex that is N- and O- glycosylated. The O-linked glycosylation is necessary to target the protein to the apical membrane. In addition, there is a segment that is both O-linked glycosylated and Ser/Thr-rich.
Sucrase-isomaltase is composed of duplicated catalytic domains, N- and C-terminal. Each domain displays overlapping specificities. Scientists have discovered the crystal structure for N-terminal human sucrase-isomaltase (ntSI) in apo form to 3.2 Å and in complex with the inhibitor kotalanol to 2.15 Å resolution.
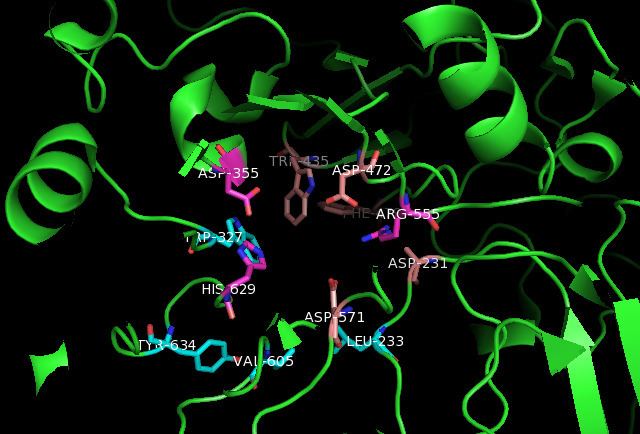
The crystal structure shows that sucrase-isomaltase exists as a monomer. The researchers claim that the observance of SI dimers is dependent on experimental conditions. ntSI’s four monomers, A, B, C, and D are included in the crystal asymmetric unit and have identical active sites. The active site is composed of a shallow-substrate binding pocket including -1 and +1 subsites. The non-reducing end of substrates binds to the pocket. While the non-reducing sugar ring has interactions with the buried -1 subsite, the reducing ring has interactions with the surface exposed +1 subsite.
The interactions between the active site of sucrase-isomaltase and the following compounds have been identified:
Currently, there are no crystal structures of ntSI in complex with an α-1,6-linked substrate or inhibitor analogue. In order to predict isomaltose binding in sucrase-isomaltase structure, a model was produced by hand. Within the -1 subsite, isomaltose’s non-reducing glucose ring was aligned to that of acarbose.
Not only has the structure of human sucrase-isomaltase been studied, but also sucrase-isomaltase’s structure in sea lions and pigs have also been analyzed.
Disease relevance
A deficiency is responsible for sucrose intolerance. Congenital sucrase-isomaltase deficiency (CSID), also called genetic sucrase-isomaltase deficiency (GSID), and sucrose intolerance, is a genetic, intestional disorder that is caused by a reduction or absence of sucrase and isomaltase Explanations for GSID include:
Furthermore, a relationship between mutations in sucrase-isomaltase and chronic lymphocytic leukemia (CLL) has been identified. These mutations cause a loss of enzyme function by blocking the biosynthesis of SI at the cell surface.
