Development status Active Operating system Available in English | Platform Linux, OSX | |
 | ||
Stable release 16.04 / 18 May 2016 (2016-05-18) | ||
Galaxy is a scientific workflow, data integration, and data and analysis persistence and publishing platform that aims to make computational biology accessible to research scientists that do not have computer programming experience. Although it was initially developed for genomics research, it is largely domain agnostic and is now used as a general bioinformatics workflow management system.
Contents
Functionality
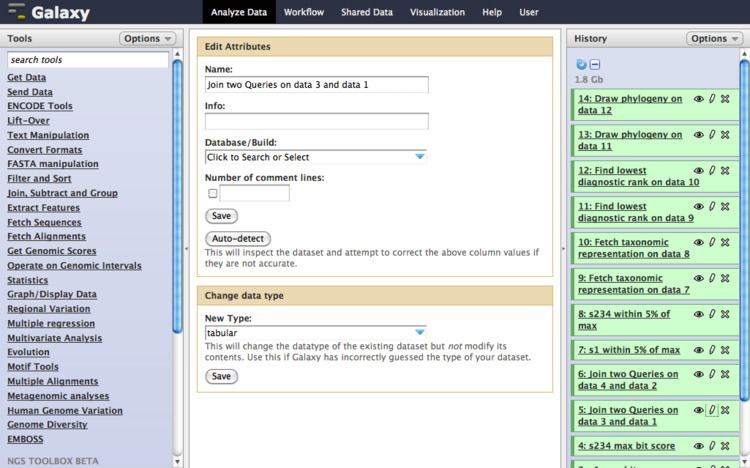
Galaxy is a scientific workflow system. These systems provide a means to build multi-step computational analyses akin to a recipe. They typically provide a graphical user interface for specifying what data to operate on, what steps to take, and what order to do them in.
Galaxy is also a data integration platform for biological data. It supports data uploads from the user's computer, by URL, and directly from many online resources (such as the UCSC Genome Browser, BioMart and InterMine). Galaxy supports a range of widely used biological data formats, and translation between those formats. Galaxy provides a web interface to many text manipulation utilities, enabling researchers to do their own custom reformatting and manipulation without having to do any programming. Galaxy includes interval manipulation utilities for doing set theoretic operations (e.g. intersection, union, ...) on intervals. Many biological file formats include genomic interval data (a frame of reference, e.g., chromosome or contig name, and start and stop positions), allowing these data to be integrated.
Galaxy was originally written for biological data analysis, particularly genomics. The set of available tools has been greatly expanded over the years and Galaxy is now also used for gene expression, genome assembly, proteomics, epigenomics, transcriptomics and host of other disciplines in the life sciences. The platform itself is actually domain agnostic and can be applied, in theory, to any scientific domain. For example, Galaxy servers exist for image analysis, computational chemistry and drug design, cosmology, climate modeling, social science, and linguistics.
Finally, Galaxy also supports data and analysis persistence and publishing. See Reproducibility and Transparency below.
Project Goals
Galaxy is "an open, web-based platform for performing accessible, reproducible, and transparent genomic science."
Accessibility
Computational biology is a specialized domain that often requires knowledge of computer programming. Galaxy aims to give biomedical researchers access to computational biology without also requiring them to understand computer programming. Galaxy does this by stressing a simple user interface over the ability to build complex workflows. This design choice makes it relatively easy to build typical analyses, but more difficult to build complex workflows that include, for example, looping constructs. (See Apache Taverna for an example of a data-driven workflow system that supports looping.)
Reproducibility
Reproducibility is a key goal of science: When scientific results are published the publications should include enough information that others can repeat the experiment and get the same results. There have been many recent efforts to extend this goal from the bench (the "wet lab") to computational experiments (the "dry lab") as well. This has proved to be a more difficult task than initially expected.
Galaxy supports reproducibility by capturing sufficient information about every step in a computational analysis, so that the analysis can be repeated, exactly, at any point in the future. This includes keeping track of all input, intermediate, and final datasets, as well as the parameters provided to, and the order of each step of the analysis.
Transparency
Galaxy supports transparency in scientific research by enabling researchers to share any of their Galaxy Objects either publicly, or with specific individuals. Shared items can be examined in detail, rerun at will and copied and modified to test hypotheses.
Galaxy Objects: Histories, Workflows, Datasets and Pages
Galaxy objects are anything that can be saved, persisted, and shared in Galaxy:
Availability
Galaxy is available:
- As a free public web server, supported by the Galaxy Project. This server includes many bioinformatics tools that are widely useful in many areas of genomics research. Users can create logins, and save histories, workflows, and datasets on the server. These saved items can also be shared with others.
- As open-source software that can be downloaded, installed and customized to address specific needs. Galaxy can be installed locally or using a computing cloud.
- Public web servers hosted by other organizations. Several organizations with their own Galaxy installation have also opted to make those servers available to others.
- As part of the GenomeSpace initiative.
Implementation
Galaxy is open-source software implemented using the Python programming language. It is developed by the Galaxy team at Penn State and Johns Hopkins University, and the Galaxy Community.
Galaxy is extensible, as new command line tools can be integrated and shared within the Galaxy ToolShed.
An example of extending Galaxy is Galaxy-P from the University of Minnesota Supercomputing Institute, which is customized as a data analysis platform for mass spectrometry-based proteomics.
Community
Galaxy is an open source project and the community includes users, organizations that install their own instance, Galaxy developers, and bioinformatics tool developers. The Galaxy project has mailing lists, a community wiki, and annual meetings.
