Group Group IV ((+)ssRNA) | ||
 | ||
Species Cassava brown streak virus | ||
General Introduction
Cassava brown streak virus (CBSV) was first described in the former Tanganyika territory of East Africa, now Tanzania 70 years ago. An endemic viral outbreak progressed throughout the Eastern African coastal cassava-growing areas from Southern Kenya, through Tanzania to the Zambezi river in Mozambique. Cassava, a tuberous, starch-rich plant (Manihot esculenta) has remained a staple food crop among countries of sub-Saharan Africa for many years, and is a primary food source for many families. Loss of such crops has a large effect on food sources and economic stability;however, the extent of the loss cannot be fully determined until the crop is harvested. Transmission is most commonly attributed to whiteflies (two whitefly species, Bemisia afer (Priesner & Hosny) and Bemisia tabaci (Gennadius)). Disease spread can be rapid, resulting in incidences exceeding 50% in some coastal regions of Tanzania and Mozambique. CBSV and other related virus strains are responsible for up to $100 million USD in losses each year in Africa due to crop destruction.
Contents
- General Introduction
- Viral classification
- Structure and Genome
- Replication Cycle
- Transmission
- Associated diseases
- Recent outbreaks
- Prevention and Antiviral Strategies
- References
Viral classification
Cassava brown streak virus belongs to the genus Ipomovirus, and the family Potyviridae. As the largest of the 34 plant virus groups and families currently recognized, the Potyviridae family contains at least 180 definitive members responsible for significant losses in agricultural, pastoral, horticultural and ornamental crops. These viruses are unique in their induction of pinwheel, or scroll-shaped inclusion bodies in the cytoplasm of infected cells. Cylindrical inclusion bodies included aggregations of virus-encoded helicase proteins. These inclusion bodies are thought to be sites of viral replication and assembly, making then an important factor in the viral lifecycle.
Structure and Genome
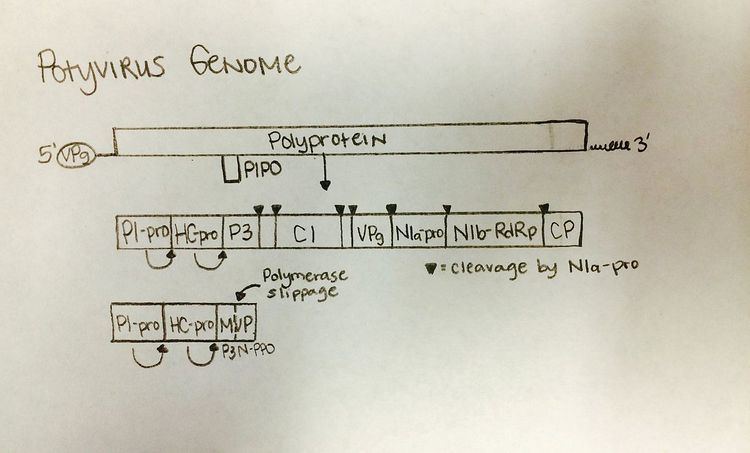
As with all of the members of the Potyviridae family, the Cassava brown streak virus has a positive, single strand RNA genome. Structurally, viruses in this family are linear, non-segmented, possess helical symmetry. The CBSV genome was measured to be between 8,900 and 10,818 nucleotides in length. This is characteristic of the average genome length among potyviridae which can be anywhere from 9.3-10.8 kb. Genetic diversity on the nucleotide level among CBSV virus isolates can be anywhere from 79.3-95.5%, showing that the virus has greater genetic diversity than other similar potyviridae or CBSV strains. The Potyvirus genome is monopartite, meaning the genome is made up of a single molecule of nucleic acid, not broken up into multiple pieces.
CBSV is believed to be a non-enveloped virus with a capsid protein coat that is most similar to that of the ipomovirus, SPMMV (Sweet potato mild mottle virus). Similar to SPMMV, CBSV was found to have larger than average coat proteins of approximately 45 kDa in size.
Replication Cycle
Upon viral entry into the host cell, the virus uncoats its outer protein shell and releases the viral RNA into the host cell's cytoplasm. CBSV is a single stranded RNA (ssRNA(+)) virus, therefore the virus must use the host cell machinery to translate the ssRNA(+) to a polyprotein which then can be processed by the viral proteases to produce RNA dependent RNA polymerase (RdRp) and other structural proteins. The viral replication happens in the cytoplasm, where a dsRNA is made from the genomic ssRNA(+). Next, the dsRNA is replicated to provide mRNA and new ssRNA(+) which can then either pass through the cycle again to be replicated, or be used later on. With this newly produced mRNA/ssRNA(+), the virus can then assemble in the cytoplasm. The protein P3N-PIPO, a movement protein, may help to initiate cell-to-cell movement and transfer.
Transmission
Potyviruses are known to be transmitted by mites and aphids. Past research has suggested that CBSV is transmitted by insects, specifically the whitefly. Scientists have used two specific species of whitefly, Bemisia afer (Priesner & Hosny) and B. tabaci (Gennadius), to experimentally test transmission rates. B. tabaci was found to be a more successful transmitter of the virus. Infection rates have been shown to correlate with the peak of whitefly population in the months of April and May. From 2002 to 2003, low numbers of whiteflies were observed in Tanzania, as well as little CBSD recorded. Likewise, in February and March 2004 when whitefly numbers soared, rapid spread of CBSD occurred. Laboratory studies, however, suggest that CBSV is transmitted poorly by B. tabaci and indicates high rates of infection in farming fields can be attributed to cutting-borne pathogens.
Associated disease(s)
Cassava brown streak virus, as a potyvirus, promotes the development of disease and other ill effects within plant life, specifically in the woody Cassava shrub in particular. Both Cassava brown streak virus and the similar Ugandan cassava brown streak virus (UCBSV), lead to the development of Cassava Brown Streak Disease (CBSD) within Cassava plants. It is also fairly common in infected plants to discover that they have been co-infected by both CBSV and UCBSV. This disease is the main source responsible for the loss of Cassava crops in sub-Saharan Africa.
The disease manifests itself in many ways throughout the plant structure. The most common symptom of this disease is necrotic local lesions present in multiple places throughout the plant. In the storage tissues of the tuberous roots, the necrotic lesions appear as brown and dry spots on the plant. Plants affected by CBSD may also develop symptoms in their leaves. CBSD is normally observed in mature or nearly mature leaves, yet does not affect expanding or immature leaves. These symptoms are often associated with the minor veins of leaves, including a type of feathery necrosis along the veins, as well as a blotchy yellow chlorosis. Symptoms vary depending on severity of infection, and in the most severe cases the lesions may cause death of dormant axillary buds. Death of these buds is likely to cause death of the entire branch of the plant.
Recent outbreaks
The most recent activity of this virus has not been expressly recorded in great detail. The most recent records of its activity show it continuing to cause widespread infection along areas of the eastern coast of Africa. These areas along the Indian Ocean coast of Africa include many of the areas where infections have persisted over time in sub-Saharan Africa such as Tanzania and Kenya. The most recent cases have shown a shift from the low altitude pattern of infection of areas below 1,000 meters above sea level. The most recent infections have branched into Uganda and the Democratic Republic of Congo where altitude levels are between 1,200 and 1,500 meters above sea level. These new areas of infection were not previously considered to be at risk.
Prevention and Antiviral Strategies
Evidence has been collected that suggests that cross-protection against a diverse range of CBSD-causing virus isolates can be achieved through RNAi silencing. The conserved CP gene, which serves multiple functions in the CBSV life cycle, was used as the transgene of interest. Three distinct RNAi constructs were designed to target the FL, N-terminal, and C-terminal of the UCBSV-CP gene: the FL-CP construct exhibiting the highest level of protection. In conclusion, the study found a significant positive correlation between the presence of siRNAs and viral protection against UCBSV. A negative drawback to this antiviral approach is that RNAi is highly sequence specific. Viruses with more than 10% nucleotide diversity require the development of an RNAi construct with multiple fragments from different virus species. A construct made out of such large sections of genes could be very unstable during cloning and transformation procedures. However, the study included two transgenic plant lines in which remained completely immune to viral infection against six unique isolates of both UCBSV and CBSV. RNAi FL-CP transgenic cassava lines also experienced 100% immunity when grafted with UCBSV infected cassava samples compared to nontransgenic control lines which became 100% infected with UCBSV over the 60-day period. In the fields, it is common for asymptomatic plants to carry the virus and serve as a source of transmission with the help of whitefly vectors or stem cuttings. However, the study also collected RT-PCR data that suggested all asymptomatic plants also lacked detectable levels of UCBSV.
