 | ||
Gene music using protein sequence of alkbh3 alkb homolog 3 alpha ketoglutarate dependent dioxygen
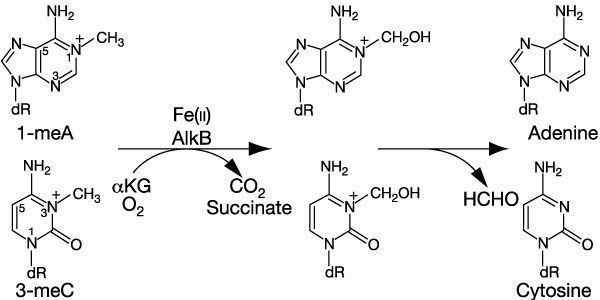
AlkB protein is a protein induced during an adaptive response and is involved in the direct reversal of alkylation damage. AlkB specifically removes alkylation damage to single stranded (SS) DNA caused by SN2 type of chemical agents. It efficiently removes methyl groups from 1-methyl adenines, 3-methyl cytosines in SS DNA. AlkB belongs to the Fe (II)/2-oxoglutarate-dependent dioxygenase superfamily and oxidatively demethylates the DNA substrate. Demethylation by AlkB is accompanied with release of CO2, succinate and formaldehyde.
Contents
- Gene music using protein sequence of alkbh3 alkb homolog 3 alpha ketoglutarate dependent dioxygen
- Gene music using protein sequence of alkbh2 alkb homolog 2 alpha ketoglutarate dependent dioxygen
- Human homologs
- Functions
- References
Gene music using protein sequence of alkbh2 alkb homolog 2 alpha ketoglutarate dependent dioxygen
Human homologs
There are nine human homologs of AlkB. They are:
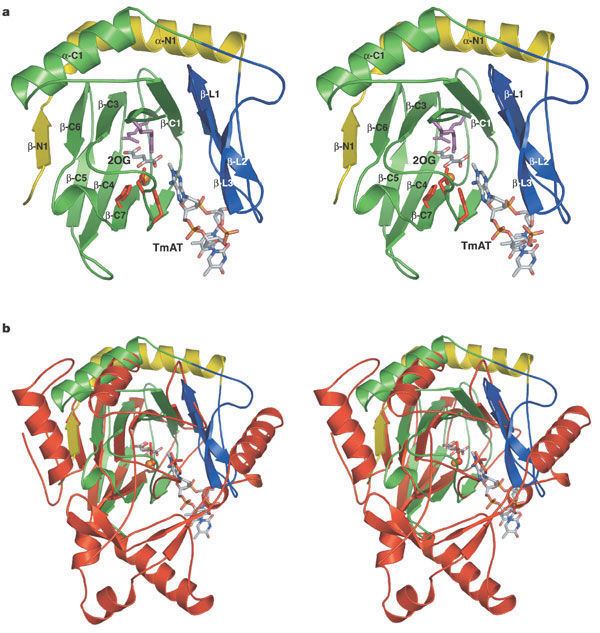
ABH3, like E. coli AlkB, is specific for SS DNA and RNA whereas ABH2 has higher affinity for damages in double-stranded DNA.
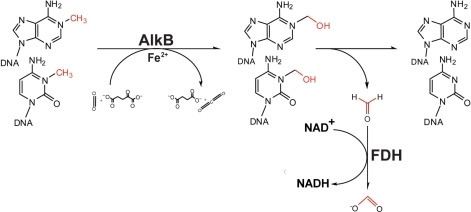
ALKBH8 has a RNA recognition motif, a methyltransferase domain, and an AlkB-like domain. The methyltransferase domain generates the wobble nucleoside 5-methoxycarbonylmethyluridine (mcm5U) from its precursor 5-carboxymethyluridine (cm5U). The AlkB-like domain generates (S)-5-methoxycarbonylhydroxymethyluridine (mchm5U)in Gly-tRNA-UCC.
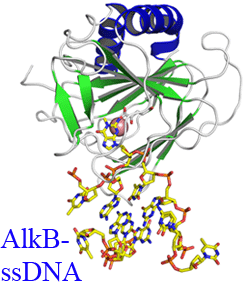
FTO, which is associated with obesity in humans, is the first identified RNA demethylase. It demethylates N6-methyladenosine in mRNA.
There is also another very different protein called AlkB or alkane hydroxylase. It is the catalytic subunit of a non-heme diiron protein, catalyzing the hydroxylation of alkanes, in aerobic bacteria that are able to utilize alkanes as a carbon source.
Functions
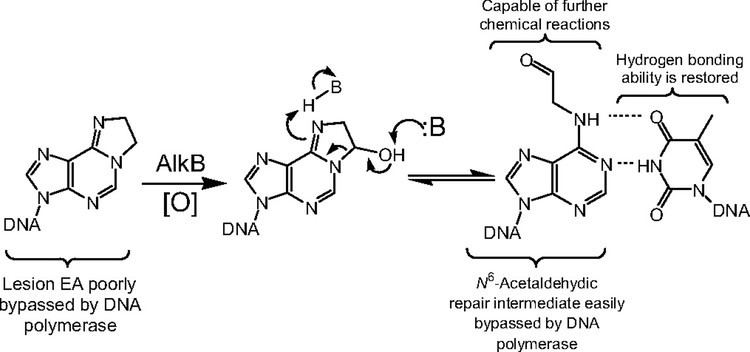
AlkB has since been shown to have an ever expanding range of substrates since its initial discovery by Sedgwick, Lindahl, Seeberg and Falnes. Not only does it remove alkylation damage from the positively charged 1-methyl adenines and 3-methyl cytosines, but also from the neutral bases of 1-methyl guanine and 3-methyl thymine. AlkB has been shown as the first example of a DNA repair enzyme converting one type of DNA damage that blocks DNA replication, to another type of damage that the DNA polymerase can traverse with ease. This was seen for the cyclic lesion ethanoadenine (not to be confused with ethenoadenine...see below), which upon hydroxylation by AlkB, affords an N6-acetaldehyde lesion, thus affording an 'adenine' hydrogen-bonding face. In contrast to the previous types of alkylation damage removed by AlkB via a hydroxylation mechanism, AlkB has been shown to epoxidize the double bond of ethenoadenine, which is hydrolyzed to a diol, and ultimately released as the dialdehyde glyoxal, thus restoring the undamaged adenine in the DNA.
