Symbol 7SK SO 0000274 | Rfam RF00100 Domain(s) Eukaryota | |
 | ||
In molecular biology 7SK is an abundant small nuclear RNA found in metazoans. It plays a role in regulating transcription by controlling the positive transcription elongation factor P-TEFb. 7SK is found in a small nuclear ribonucleoprotein complex (snRNP) with a number of other proteins that regulate the stability and function of the complex.
Contents
Structure
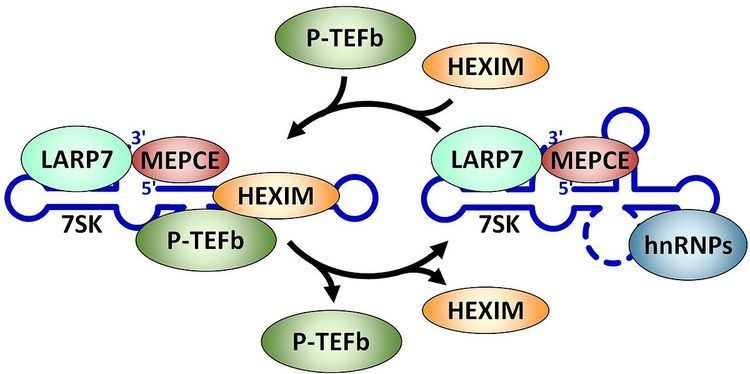
An early study indicated that 7SK in cells is associated with a number of proteins and probing of the secondary structure suggested a model for base pairing between different regions of the RNA. A breakthrough in the function of the 7SK snRNP came with the finding that the positive transcription elongation factor P-TEFb was a component of the complex. 7SK associates with and inhibits the cyclin dependent kinase activity of P-TEFb through the action of the RNA binding proteins HEXIM1 or HEXIM2. The gamma phosphate at the 5' end of 7SK is methylated by the methylphosphate capping enzyme MEPCE which is a constitutive component of the 7SK snRNP. A La related protein LARP7 is also found associated with 7SK, presumably in part through its interaction with the 3' end of the RNA. Reduction of either MEPCE or LARP7 by siRNA mediated knockdown leads to destabilization of 7SK in vivo. A subset of 7SK snRNPs lack P-TEFb and HEXIM, but contains hnRNPs instead.
Function
The major function of the 7SK snRNP is control of the P-TEFb, a factor that regulates the elongation phase of transcription. The kinase activity of P-TEFb is inhibited when the factor is in the 7SK snRNP. P-TEFb can be released from the 7SK snRNP by either the HIV transactivator Tat or the bromodomain containing protein BRD4. This release leads to a conformational change in 7SK RNA and the ejection of HEXIM. hnRNPs stabilize the complex lacking P-TEFb and HEXIM. After P-TEFb functions on specific genes it is re-sequestered in the 7SK snRNP by an unknown mechanism. The 7SK snRNP has been characterized in both human and Drosophila. Detailed review.
