 | ||
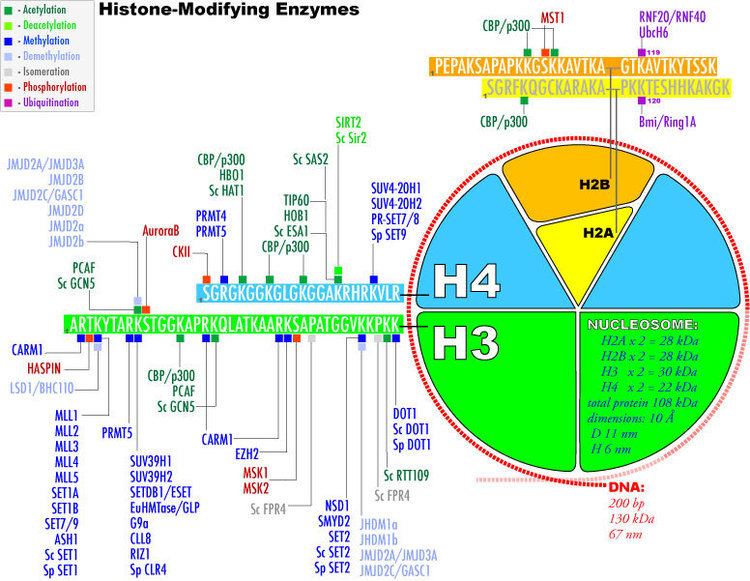
The packaging of the eukaryotic genome into highly condensed chromatin makes it inaccessible to the factors required for gene transcription, DNA replication, recombination and repair. Eukaryotes have developed intricate mechanisms to overcome this repressive barrier imposed by the chromatin. The nucleosome is composed of an octamer of the four core histones (H3, H4, H2A, H2B) around which 147 base pairs of DNA are wrapped. Several distinct classes of enzyme can modify histones at multiple sites. The figure on the right enlists those histone-modifying enzymes whose specificity has been determined. There are at least eight distinct types of modifications found on histones (see the legend box on the top left of the figure). Enzymes have been identified for acetylation, methylation, demethylation, phosphorylation, ubiquitination, sumoylation, ADP-ribosylation, deimination, and proline isomerization. For a detailed example of histone modifications in transcription regulation see RNA polymerase control by chromatin structure and table Influence of modifications on gene expression in mammalian cells.
