 | ||
An ancestry-informative marker (AIM) is a set of polymorphisms for a locus, generally from humans, which exhibits substantially different frequencies between populations from different geographical regions.
By using a number of AIMs one can estimate the geographical origins of the ancestors of an individual and ascertain what proportion of ancestry is derived from each geographical region. By using a suite of these markers more or less evenly spaced across the genome, they can be used in a cost-effective way to discover novel genes underlying complex diseases in a technique called admixture mapping or mapping by admixture linkage disequilibrium.
There are an estimated 15 million SNP (Single-nucleotide polymorphism) sites (out of roughly 3 billion base pairs, or about 0.4%) from among which AIMs may potentially be selected.
As one example, the Duffy Null allele (FY*0) has a frequency of almost 100% of Sub-Saharan Africans, but occurs very infrequently in populations outside of this region. A person having this allele is thus more likely to have Sub-Saharan African ancestors.
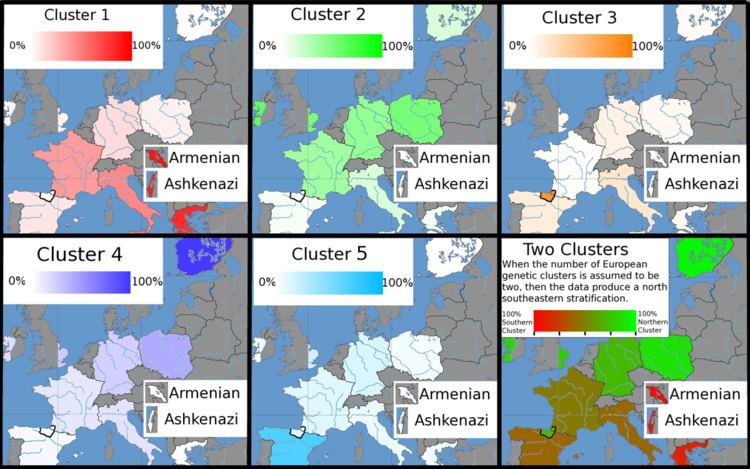
Collections of AIMs have been developed that can estimate the geographical origins of ancestors from within Europe.
